Press-room / news / Science news /
The new method for T-cell receptor alpha chain clonality assessment can be used for minimal disease monitoring in leukemia
Monitoring of minimum residual disease (MRD) is one of the most important diagnostic tests in the treatment of acute and chronic lymphoblastic leukemia. Currently, several methods for MRD are used, the most sensitive of which is the assessment and monitoring of clonal rearrangements in the immunoglobulin genes characteristic of tumor cells. High sensitivity, up to 1 tumor cell per million, is achieved by using next generation highthroughput sequencing technology (Illumina).
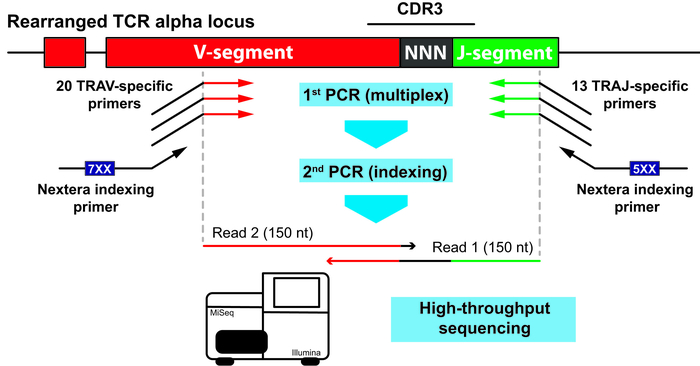
Until recently, 6 genes of the immunoglobulin receptor superfamily (IGH, IGK, IGL, TRB, TRD, and TRG) were used to monitor MRD, while the TRA gene encoding for T-cell receptor alpha chain was not utilized. A group of scientists from Laboratory of Comparative and Functional Genomics in collaboration with colleagues from Masaryk universities and Dmitry Rogachaev center for pediatric hematology, oncology and immunology developed a technique to identify and monitor clonal rearrangements in this last immunoglobulin gene. This approach is based on a combination of multiplex PCR, highthroughput sequencing and an original data analysis protocol that allows to correct the quantitative bias that occurs during amplification. The method was tested on 112 samples of patients with acute lymphoblastic leukemia or lymphoma and it was shown that about half of the samples have characteristic rearrangements of the TRA gene. The inclusion of a new gene into the MRD monitoring protocol will help to significantly increase its reliability and to reduce the number of patients for whom it was not possible to detect at least one characteristic rearrangement. The developed method can also be used to analyze the repertoires of T-cell receptors in samples for which RNA is not available. Results of the study are published in British Journal of Haematology.
december 27, 2019

