Press-room / news / Science news /
Prophage-derived regions in Curtobacterium genomes: good things, small packages
Curtobacterium is a genus of Gram-positive bacteria within the order Actinomycetales. Some Curtobacterium species are harmful pathogens of agricultural crops such as soybean, dry beans, peas, sugar beet and beetroot, which occur throughout the world. About 200 publicly available genomes of Curtobacterium species, including environmental metagenomic sequences, were inspected for the presence of sequences of possible prophage origin using bioinformatic methods.
One of the predicted temperate phages was induced from the Curtobacterium genome. Bioinformatic analysis of the modelled proteins encoded in prophage-derived regions led to the discovery of some 100 putative glycopolymer-degrading enzymes. These proteins can be considered for the experimental design of new antibacterials against Curtobacterium phytopathogens.
The results are published in the International Journal of Molecualr Sciences.

Figure 1. Simplified genetic maps of the representatives of Curtobacterium prophages. Arrows indicate the
direction of transcription.
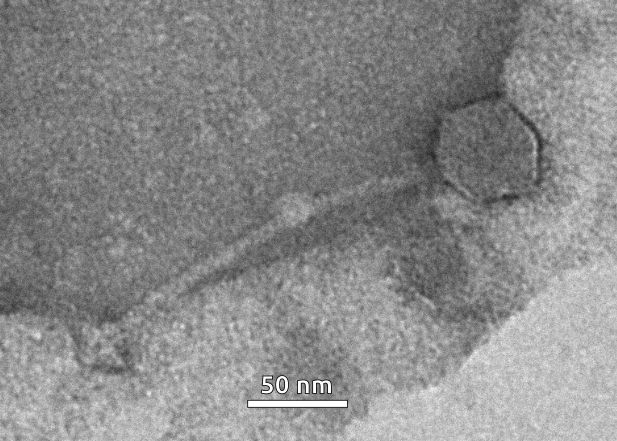

Figure 2. Electron microscopy image of phage particle induced by mitomycin C from the bacterial culture of
Curtobacterium sp. VKM Ac-2884. The scale bar is 50 nm.
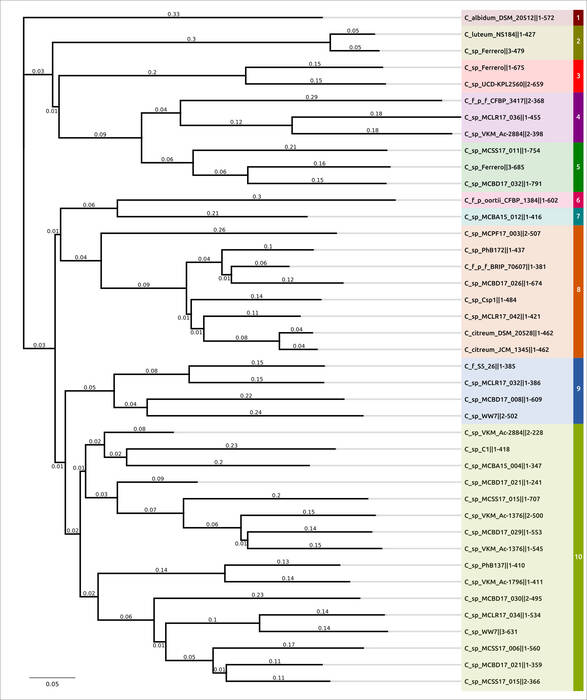
Figure 3. Phylogenetic tree based on amino acid sequences of glycopolymer-degrading enzymes using structural
similarity assessed with TM-scores. Branches belonging to different clusters are highlighted using colour.
The last three digits in the names of structures indicate the length of protein sequences.
january 23, 2023


