Лаборатория химии метаболических путей
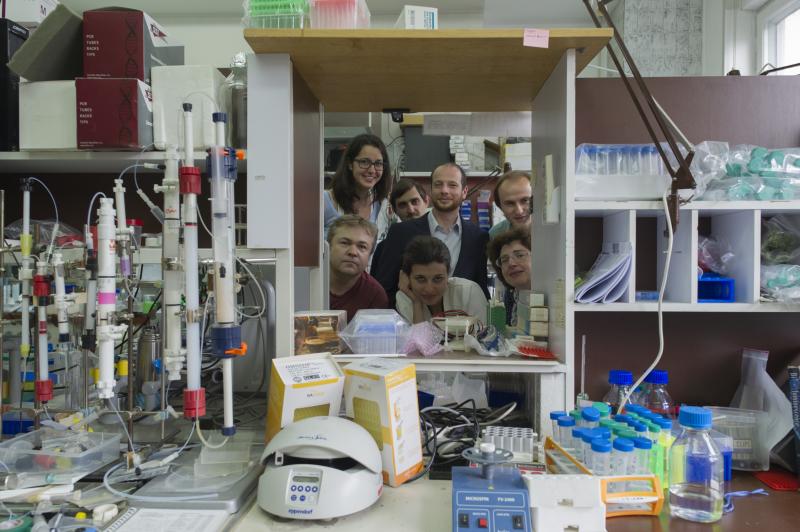
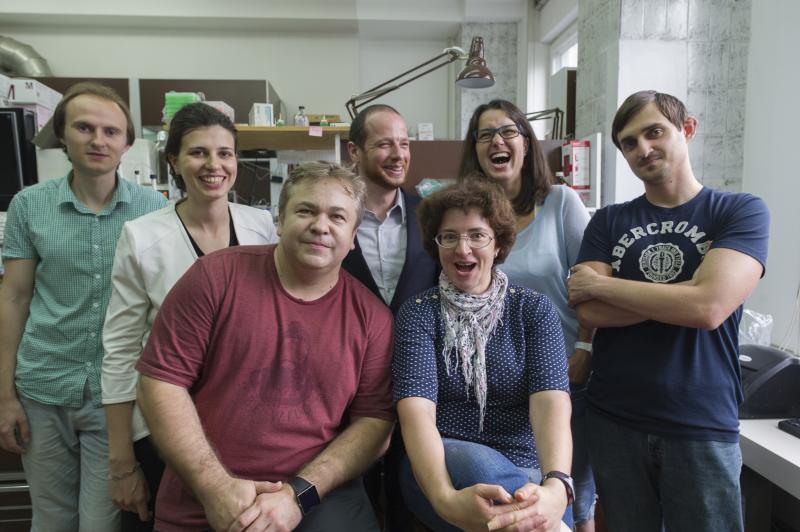
Лаборатория создана в 2017 году на основе Группы синтеза природных соединений, которая, в свою очередь, работала с 2002 года при Лаборатории молекулярных технологий ИБХ РАН, возглавляемой академиком Сергеем Анатольевичем Лукьяновым. Основным направлением исследований группы является применение методов органического синтеза для решения актуальных проблем в биохимии, молекулярной биологии и медицинской химии.
Деятельность группы включает в себя:
- структурный дизайн и синтез модельных соединений для изучения биохимических процессов
- полный синтез природных соединений
- дизайн, получение и тестирование лекарственных средств
- проведение совместных биомедицинских и биологических исследований
 |  |
 | |

- Изучение механизма биолюминесценции высших грибов

Фотография профессора Кассиуса Стевани (Университет Сан-Пауло, Бразилия)
Статья в The Guardian о нашей работе
- Исследование биолюминесценции морского червя Chaetopterus variopedatus
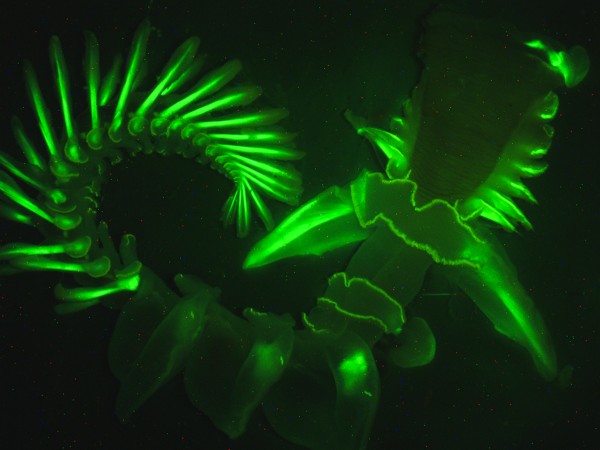
Фотография Дмитрия Дейна, Scripps Institution of Oceanography at UC San Diego
- Изучение механизма биолюминесценции сибирского почвенного червя Fridericia heliota
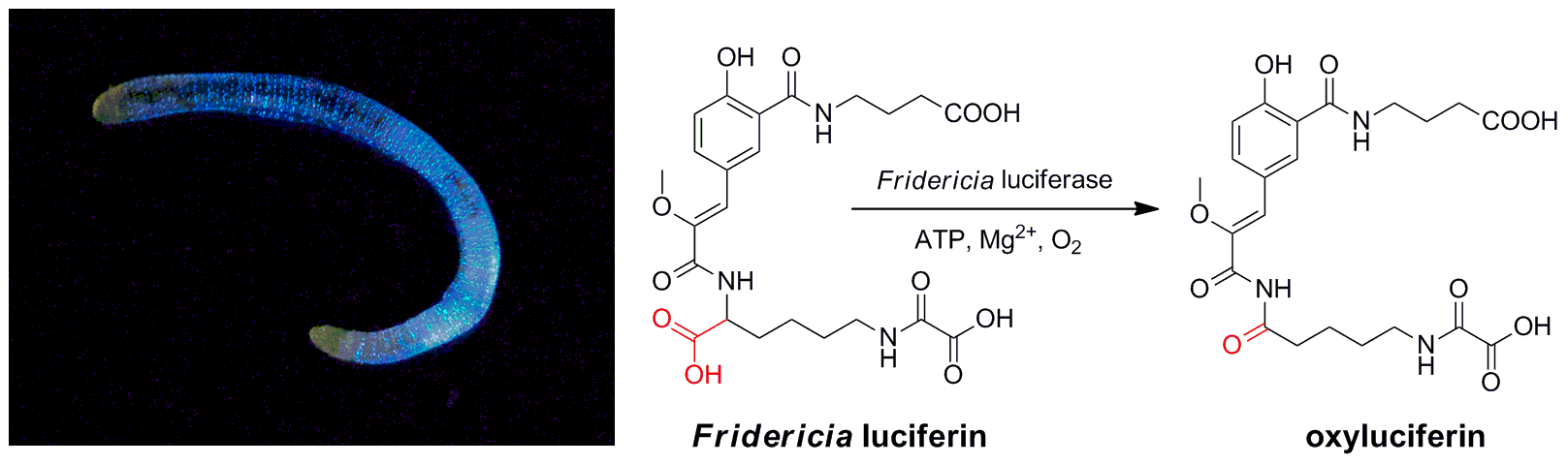
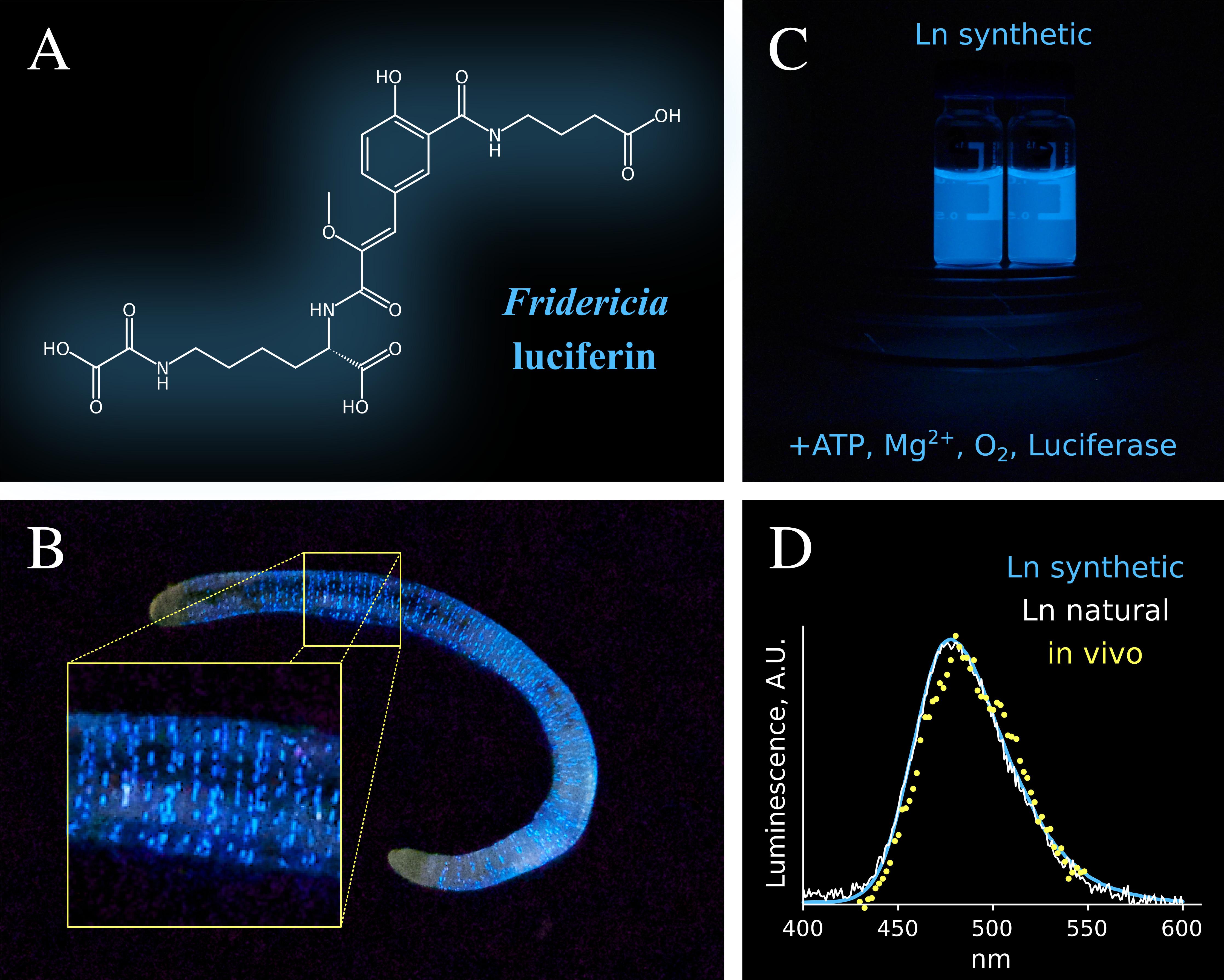
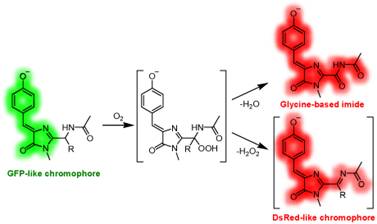
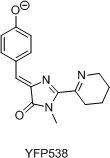
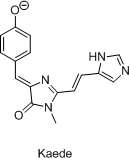
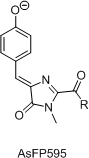
- Разработка нового класса флюоресцентных красителей на основе хромофора GFP
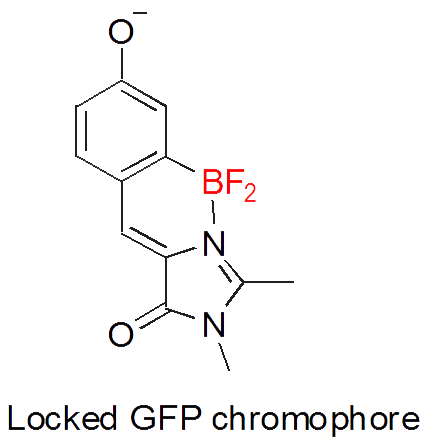
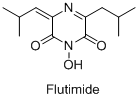
- Разработка противовирусных лекарственных препаратов - структурных аналогов Флутимида
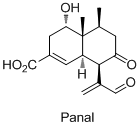
- Полный энантиоселективный синтез грибкового терпеноида Паналя из биолюминесцентных грибов Panellus stipticus

- Изучение механизма биолюминесценции светящихся грибов

- Установление биосинтеза морских люциферинов: целентеразина и Cypridina
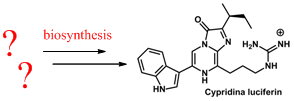
- Разработка флуорогенных датчиков для важных белковых структур







 Загрузка...
Загрузка...Научные проекты
 Загрузка...
Загрузка...Ямпольский Илья Викторович
Москва, ул. Миклухо-Маклая, 16/10 — На карте
ORCID: 0000-0003-2558-2476, Scopus: 7801466424
52 корпус, 356 комната
+7 (499) 724-84-77
 Загрузка...
Загрузка...
