Пресс-центр / новости / Наука /
Бактериофаг Pseudomonas MD8: генетический мозаицизм и проблемы таксономической классификации лямбдоидных бактериофагов
Фундаментальные вопросы эволюции вирусных геномов - важнейшая тема вирусологических исследований. В результате совместной работы вирусологов из Лаборатории молекулярной биоинженерии ИБХ и Лимнологического института РАН выявлена группа бактериофагов опасного патогена Pseudomonas, исследованы геномы этих бактериальных вирусов и показано, что на их формирование оказали большое влияние множественные горизонтальные переносы, что привело к ярко выраженному генетическому мозаицизму. Были выдвинуты гипотезы о происхождении новой группы и предложены основные принципы таксономической классификации лямбдоидных фагов.
Геномика лямбдоидных (λ-подобных) бактериофагов находится в центре внимания нескольких поколений ученых с момента открытия фага λ Эстер Ледерберг в 1949 году и последующего роста интереса к этому фагу. Идентификация генетических механизмов, ответственных за принятие решение о лизогении, процессах лизиса и адсорбции, внесли огромный вклад в понимание основных принципов природы фаговой инфекции. Фаг λ и родственные ему фаги остаются в центре исследований, посвященных молекулярным деталям инфекционного цикла фагов.
Исследования, последовавшие за открытием фага λ, выявили комплекс основных принципов организации генома этого фага и других умеренных бактериофагов. Геномные исследования продемонстрировали высокий уровень генетического мозаицизма λ-подобных фагов и одинаковую общей структуре генома. Этот генетический мозаицизм может быть вызван гомологичной рекомбинацией, происходящей при заражении клетки, несущую профаг с соответствующими гомологиями, или при неизбирательной негомологичной рекомбинацией с последующим отбором на функциональные фаги. Умеренный образ жизни лямбдоидных фагов и процессы рекомбинации этих фагов играют значительную роль в бактериальной эволюции, включая адаптацию и геномную диверсификацию бактериальных патогенов.
Pseudomonas MD8 (Рисунок 1) – умеренный фаг с λ-подобным геномом, который был получен из пресной воды озера Байкал в 2005 и 2010 годах. Фаг инфицирует Pseudomonas aeruginosa, условно-патогенный микроорганизм, который может вызывать хронические инфекции, приводящие к значительной заболеваемости и смертности. Недавние исследования обнаружили новые бактериофаги умеренного климата, заражающие P. aeruginosa. Предварительный сравнительный биоинформатический анализ генома фага MD8 показал наличие как генов, относящихся к ранее описанным лямбдоидным фагам, так и генов, специфичных к MD8 или другим фагам Pseudomonas, включая возможные факторы вирулентности. Анализ также выявил проблемы, связанные с дифференциацией и таксономической классификацией фагов Pseudomonas и их генетическим мозаицизмом.
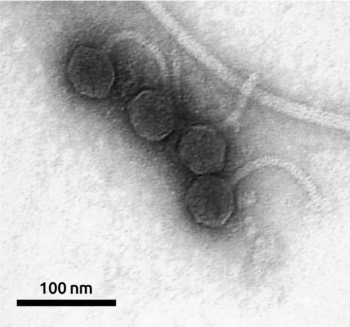
Рисунок 1. Электронная микрофотография бактериофага MD8. Шкала 100 нм
Анализ фаговых геномов выявил группу других фагов Pseudomonas, связанных с фагом MD8, а геномная структура MD8-подобных фагов указывает на обширный обмен генами, включающий даже самые консервативные белки и ведущий к высокой степени мозаицизма фагового генома (Рисунок 2). Геномы MD8-подобных фагов кодируют относительно далекие от фага λ белки, но многие из этих белков обладают функциональностью, аналогичной белкам фага λ. Множественные горизонтальные переносы и мозаичность генома MD8, родственных фагов и других λ-подобных фагов вызывают вопросы о принципах таксономической классификации представителей этой обширной группы фагов.
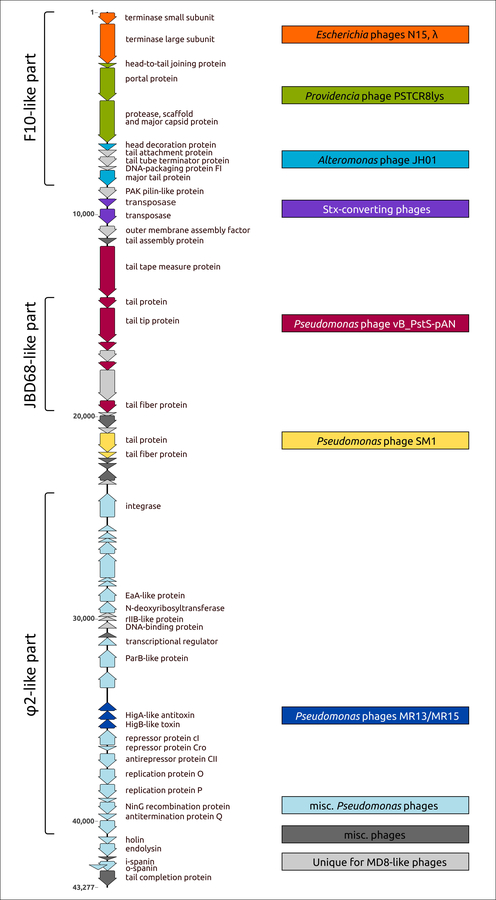
Рисунок 2. Генетическая карта фага Pseudomonas MD8, демонстрирующая расположение геномных модулей, идентифицированных путем сравнения последовательностей и филогенетического анализа. Части геномов, в целом более похожие на фаги F10, JBD68 и φ, принадлежащих к группе MD8, показаны в скобках слева
С момента первого опубликованного отчета в 1971 году обновление таксономической классификации вирусов было основной задачей Международного комитета по таксономии вирусов (ICTV). С 1971 г. было опубликовано несколько отчетов, содержащих результаты тщательных исследований, но несколько тысяч геномов Siphoviridae до сих пор остаются не классифицированными. Для таксономической классификации бактериофагов были разработаны различные подходы их кластеризации, но предварительные исследования MD8 и родственных фагов показали несоответствие между результатами различных методов.
В статье обсуждаются трудности эволюционного анализа и таксономической классификации MD8, родственных фагов и λ-подобных фагов в целом. В работе описаны биологические и морфологические свойства MD8, произведён поиск в базе данных связанных фагов, и родственные геномы сгруппированы с использованием различных методов. Основные гены и соответствующие белки были изучены с помощью анализа гомологичных последовательностей, а также филогенетического и структурного биоинформатического анализа. Были проанализированы найденные родственные геномы, изучена протеомная филогениия, обнаружены модули, полученные путем горизонтального переноса, и реконструированы возможные источники возникновения мозаичной картины генома MD8. В статье обсуждаются результаты выполненных исследований и их значение для таксономии лямбдоидных фагов, сделаны предложения по алгоритму таксономической классификации λ-подобных бактериофагов (Рисунок 3). Основываясь на эволюционной истории большинства белков MD8-подобных фагов и результатах геномного анализа, группу, состоящую из 26 MD8-подобных фагов, было предложено отнести к двум новым родам.
Работа опубликована в журнале International Journal of Molecular Sciences.
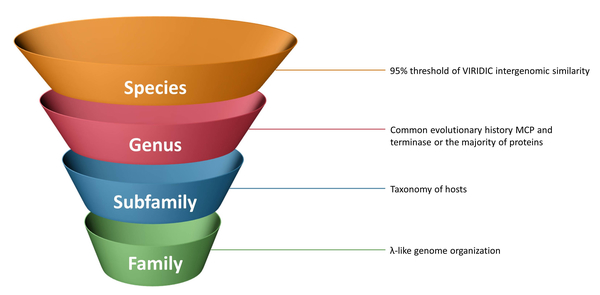
Рисунок 3. Возможные критерии таксономической классификации лямбдоидных бактериофагов
8 октября 2021 года


