Пресс-центр / новости / Наука /
Эволюция чехольных белков бактериофагов
Группа ученых из Лаборатории молекулярной биоинженерии ИБХ РАН с помощью нового программного обеспечения AlphaFold 2 смоделировала и проанализировала 112 сократительных белков оболочки хвоста фага (TShP), представляющих различные группы бактериофагов и архейных вирусов с миовирусной морфологией. Было показано, что общий домен всех исследованных белков оболочки, включая вирусные белки и белки ССТТ, включает как N-концевую, так и С-концевую части, тогда как остальные части состоят из одного или нескольких умеренно консервативных доменов, полученных в ходе эволюции бактериофагов.
Анализ экспериментально определенных и смоделированных структур свидетельствует о наличии общего консервативного ядра, имеющегося у всех анализируемых чехольных белков (ЧБ), в том числе чехольных белков бактериофагов (ЧББ), чехольных белков системы секреции VI типа (T6SS), бактериоцинов и антипитающих (anti-feeding) профаговых (AFP ) чехольных белков. Большинство ЧББ, исследованных в работы, отличались от нефаговых чехольных белков наличием дополнительных доменов. Эти домены, по-видимому, расположены вдали от той части ЧББ, которая контактирует с хвостовой трубкой. Такое расположение препятствует нарушению работы сократительного механизма. Таким образом, эволюция ЧББ была обусловлена его биологической ролью. Филогенетический анализ на основе структурного сходства указывал на родство белков оболочки системы секреции VI типа и ЧББ гигантских фагов. Это может указывать на древнюю дивергенцию белков фаговой оболочки и T6SS. Кроме того, сходство между чехольными белками системы AFP, TShPs грамположительных бактерий и предсказанными ЧББ архей также может иметь древнее происхождение и быть следствием специализации AFP. Напротив, ЧБ диффоцина и пиоцина, по-видимому, возникли позже и могут быть полифилетическими. Интересно, что белок оболочки диффоцина Peptoclostridium difficile структурно ближе к фагам Clostridium, чем ко всем другим белкам оболочки, а белок оболочки пиоцина Pseudomonas aeruginosa структурно близок к ЧББ фагов, инфицирующих бета- и гаммапротеобактерии. Происхождение фагоподобных сократительных машин требует специальных исследований с участием большей репрезентативной группы.
Размер ЧББ и количество дополнительных доменов у этого белка коррелирует с размером генома, – малые фаги обладают более короткими одно- или двухдоменными ТШП. Это наблюдение кажется осмысленным, так как дополнительные домены не являются существенными для сборки и работы сократительного механизма, но потребляют ресурсы для переноса дополнительного генетического материала и синтеза белка во время инфекции. Крупные фаги часто имеют мультидоменные ЧББ. Здесь предполагается, что предковая форма фаговых ЧББ обладала одним основным доменом, а в ходе эволюции фагов, сопровождавшейся увеличением размера генома, фаговые ЧББ приобретали дополнительные домены. Этот процесс может иметь некоторые общие черты с приобретением дополнительных функциональных генов в ходе эволюции джамбо-фагов.
Необходимость расходования дополнительных ресурсов в результате наличия более крупных белков оболочки должна быть оправдана конкурентными преимуществами, обеспечиваемыми дополнительными доменами. Большинство дополнительных доменов в исследованных ЧББ имели иммуноглобулин-подобную (Ig-подобную) укладку. Можно предположить, что наличие дополнительных Ig-подобных доменов ТШП способствует адгезии фагов к бактериям. Было показано, что Ig-подобные домены являются предметом общего горизонтального обмена между различными классами как литических, так и умеренных фагов, и Ig-подобные домены могут играть вспомогательную роль в фаговой инфекции, взаимодействуя с углеводами на поверхности бактериальной клетки. Еще одна гипотеза может заключаться в том, что эти домены могут участвовать в формировании сложных хвостовых отростков, обнаруженных у некоторых крупных фагов, поскольку они часто участвуют в белок-белковых взаимодействиях, которые, в свою очередь, также способствуют клеточной адгезии. Возможно также, что наличие дополнительных доменов способствует повышению жизненно важной для фагов стабильности вириона за счет упрочнения собранного хвоста за счет взаимодействия между дополнительными доменами ЧББ.
Работа опубликована в журнале Viruses.
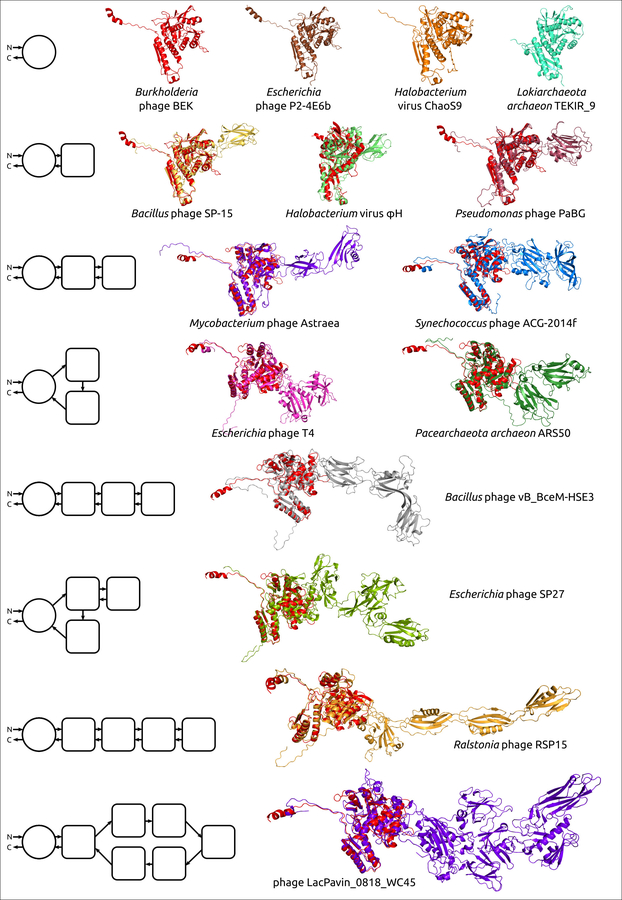
Рисунок 1. Примеры структурной архитектуры смоделированных сократительных чехольных белков бактериофагов (ЧББ), совмещённые с ЧББ фага Burkholderia (красный). Слева показана структурная архитектура белков. Основной домен изображен в виде круга, а дополнительные домены представлены в виде квадратов со скругленными углами. Направление полипептидной цепи от N-конца к С-концу показано стрелками.
16 июня 2022 года


