Пресс-центр / новости / Наука /
Использование комплексного подхода, включающего предсказания AlphaFold, для эволюционной таксономии вирусов реалма Duplodnaviria
Классификация высокоранговых таксонов, включая семейства и отряды – важная задача для таксономии вирусов реалма Duplodnaviria. В настоящем исследовании сотрудники Лаборатории молекулярной инженерии ИБХ РАН совместно с коллегами из Лимнологического института СО РАН проанализировали эволюционные отношения консервативных вирусных белков, представляющих различные вирусы, включая все классифицированные семейства Duplodnaviria.
Анализ был проведен с использованием предсказаний AlphaFold, структурных сравнений и различных филогенетических методов. Результаты анализов указывали на, в целом, высокое качество моделирования с помощью AlphaFold и возможность использования предсказаний AlphaFold, совместно с другими методами, для реконструкции эволюционных взаимоотношений между отдаленными группами вирусов.
Работа опубликована в журнале Biomolecules.
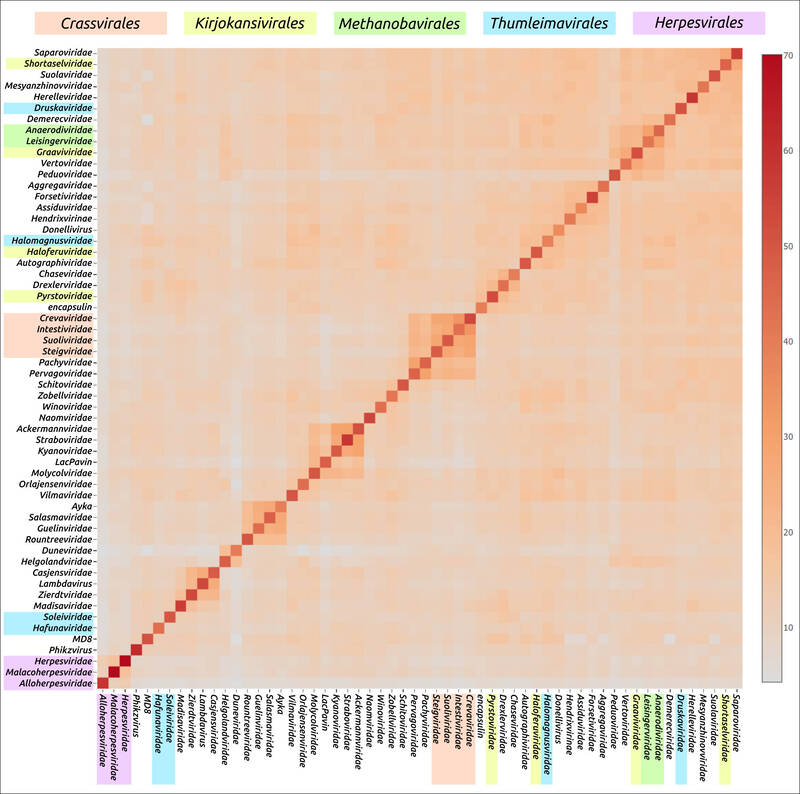
Рисунок 1. Тепловая карта, полученная с помощью сравнения 57 главных капсидных белков.
18 января 2023 года


