Пресс-центр / новости / Наука /
Фрактальные капли
Биологические полимеры (белки, нуклеиновые кислоты и их смеси) способны образовывать макромолекулярные комплексы в растворе, которые при достижении характерных концентраций насыщения полимера увеличиваются в размере и при достижение дифракционного предела (> 0.25 µm) становятся видимыми под микроскопом. Наблюдаемые сферических образования жидкой природы в зависимости от контекста имеют разные наименования: «коацерватные капли», бимолекулярные конденсаты, и др. Несмотря на то, что такие полимерные капли (в первую очередь – белковые) уже более века привлекают внимание ученых, в том числе как потенциальные «протоклетки» в «первичном бульоне» согласно одной из гипотез происхождения жизни, предложенной акад. Опариным, активное исследования их роли в функционировании живой клетки началось в последние десятилетия.
Согласно современным представлениям, бимолекулярные конденсаты активно участвуют в различных процессах внутриклеточной организации (ядрышко, тельца Кахаля, стрессовые гранулы, РНК гранулы в нейронах, и другие не мембранные органеллы клетки). В частности, было показано, что неупорядоченные белки (IDP) являются одним из ключевых компонентов во многих таких конденсатах и непосредственно участвуют в их образовании, а также способны формировать подобные капли in vitro (Banani et al., 2017; Uversky, 2021). В то же время, принципы пространственной и динамической организации IDP-конденсатов на разных уровнях, определяющие взаимосвязь между конформационными свойствами и паттернами взаимодействий белковых последовательностей и характеристиками сформированных ими конденсатов, остаются неясными. В совместной исследовательской работе ученых из ИБХ РАН и Венского университета была предложена фрактальная модель, которая позволяет численно описать архитектуру конденсата, исходя из атомистических параметры формирующих его неупорядоченных белков. Данная модель также находится в согласии с недавно предложенной концепцией образования конденсатов путем сочетания фазового разделения и перколяции (Mittag & Pappu, 2022).
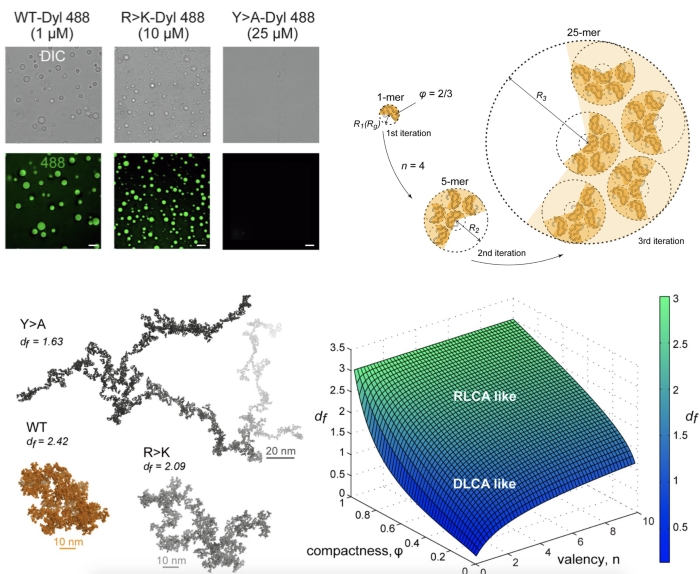
В качестве модельной системы использовали N-концевой неупорядоченный фрагмент (80 остатков) регулятора транскрипции Lge1 (Lge11-80 ) и его функциональных мутантов из дрожжей, для которых данные расчетов микросекундной молекулярной динамики (МД) для единичной молекулы и их ансамбля были сопоставлены с результатами биофизических экспериментов in vitro. В частности, предложенный формализм позволяет описывать архитектуру конденсатов в разном диапазоне размеров как функцию валентности (по аналогии с координационным числом в кристаллических решетках) и компактности формирующего их белка. При этом полученные на основании данных МД значения фрактальной размерности, описывающей архитектуру конденсатов Lge11-80 и его мутантов, соответствуют результатам экспериментов in vitro. Разработанный формализм, также позволяет получить атомистическую структуру конденсатов в микрометровом диапазоне, используя МД-конформации исходного белка. Таким образом, предложенная теоретическая модель описывает белковые конденсаты в разном диапазоне шкал в соответствии с общими принципами самоорганизации коллоидных систем. Исследование опубликовано в eLife.
Литература
- Banani et al., Nat. Rev. Mol. Cell Biol., 2017, 18(5), 285-298. doi:10.1038/nrm.2017.7
- Uversky, V. N. Annu. Rev. Biophys., 2021, 50(1), 135-156. doi:10.1146/annurev-biophys- 062920-063704
- Mittag, T., & Pappu, R. V. Mol. Cell, 2022, 82(12), 2201-2214. doi:10.1016/j.molcel.2022.05.018
11 сентября 2023 года


