Пресс-центр / новости / Наука /
Множественные паралоги регулятора развития переднего мозга – гена foxg1 - у миног и осетровых: наследие геномных дупликаций в ранней эволюции позвоночных
Передний мозг позвоночных, в том числе его уникальный отдел – конечный мозг, обеспечивают высшие формы нервной деятельности у животных и человека. В качестве одного из ведущих регуляторов раннего развития переднего мозга описан ген foxg1. Показано, что нарушения экспрессии foxg1 приводят к аномалиям развития и уменьшению размеров переднего мозга, у человека называемые синдромом FOXG1.
Ранее у большинства позвоночных и их ближайших родственников рассматривалось наличие одного гена foxg1, который часто применяется в качестве молекулярного маркера переднего мозга и связанных с ним структур.
В рассматриваемой статье в журнале Frontiers in Cell and Developmental Biology мы впервые описали наличие множественных паралогов foxg1 у миног, как представителей наиболее эволюционно древней группы позвоночных – бесчелюстных и осетровых, как одной из архаичных групп челюстноротых.
Для оценки родства генов foxg1 у миног и челюстноротых был проведен анализ их локальной геномной синтении (Рисунок 1), филогенетический анализ кодируемых ими белков и исследование особенностей экспрессии паралогов foxg1 у речной миноги и стерляди.
Было показано, что наблюдаемая у высших позвоночных (птиц, млекопитающих и некоторых рептилий) ситуация наличия одного гена foxg1 не является базовой для позвоночных. У филогенетически древних групп - хрящевых, осетровых и костных рыб присутствуют, как правило, три гена foxg1. У некоторых представителей осталось два гена (C. milii), а у костистых рыб, вследствие прошедшей у них дополнительной геномной дупликации есть четыре паралога foxg1. Поскольку у ближайших родственников позвоночных (ланцетников и оболочников) обнаружено по одному foxg1 гену, была выдвинута гипотеза о появлении трех генов foxg1 позвоночных в результате двух раундов геномных дупликаций в ранней эволюции позвоночных. Эта гипотеза была подтверждена оценкой сроков дупликации паралогов foxg1 в разных группах позвоночных.
Анализ пространственной экспрессии показал, что все три foxg1 гена миног экспрессируются в конечном мозге, хотя паттерны экспрессии генов foxg1 различаются (Рисунок 2). У стерляди в переднем мозге экспрессируется только ген foxg1a.
Обнаруженная экспрессия генов foxg1 в переднеголовных структурах, сенсорных органах и черепных нервах у бесчелюстных и челюстноротых может свидетельствовать о важной эволюционной роли генов foxg1 в формировании плана строения этих морфологических признаков позвоночных. При этом нами не обнаружено достоверной парной ортологии между отдельными паралогами foxg1 бесчелюстных и челюстноротых, которая могла бы свидетельствовать в пользу гипотезы одного или двух раундов общей полногеномной дупликации до разделения этих эволюционных линий.
Установлено, что выявленная филогения паралогов foxg1 осетровых соответствует модели предковой дупликации с последующей асинхронной редиплоилизацией, а дупликация генов foxg1b до разделения линий Polyodon и Acipenser указывает на геномную дупликацию, произошедшую на уровне их общего предка.
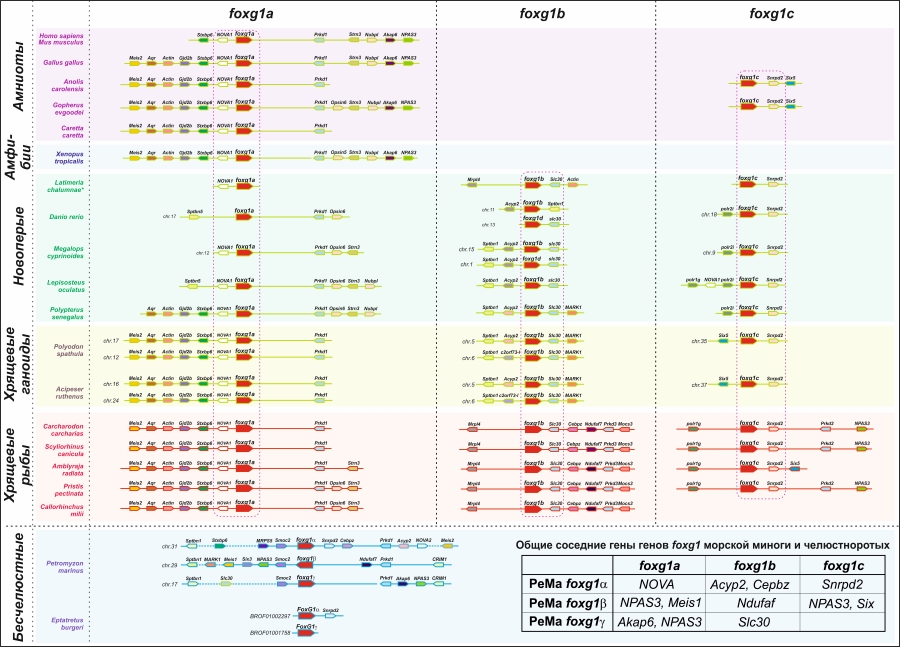

Рисунок 1. Анализ локальной геномной синтении генов foxg1 позвоночных.
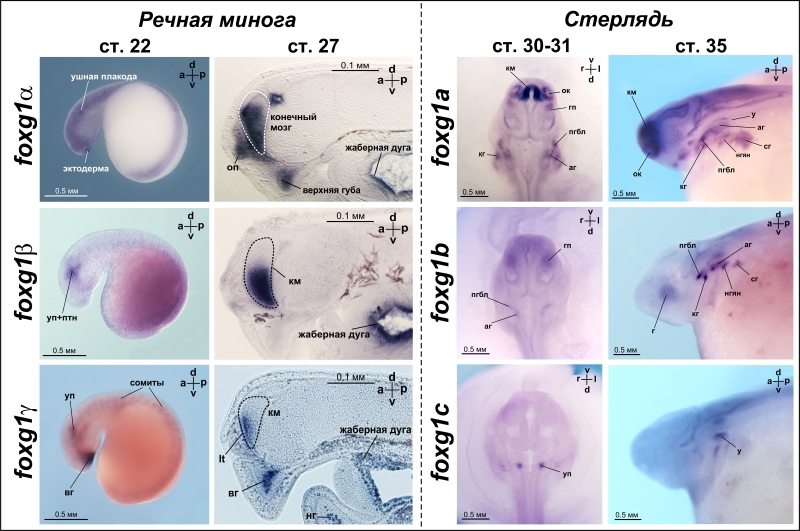
Рисунок 2. Особенности экспрессии паралогов foxg1 у речной миноги (бесчелюстные) и стерляди (осетровые, как архаичная группа челюстноротых).
25 января 2024 года


