Лаборатория молекулярных основ эмбриогенеза
Лаборатория изучает молекулярно-генетические механизмы раннего развития и эволюции нервной системы, а также регенерации больших придатков тела на моделях низших позвоночных.
Один из проектов посвящен исследованию у позвоночных животных моногенного семейства гомеобоксных генов Anf (Dev Biol, 1992, 152, 373-382; Development, 1995, 121, 3839-3847; Gene, 1997, 200, 25-34). Впервые установлено, что ген присутствует в геномах исключительно у позвоночных животных, включая человека, и отсутствует у беспозвоночных. Он контролирует развитие уникального отдела мозга позвоночных – конечного мозга – и в клетках зачатка конечного мозга играет роль репрессора транскрипции, подавляя экспрессию генов, которые усиливают формирование задних отделов мозга (Development, 1999, 126, 4513- 4523; Gene, 2002, 285, 279-286; Development, 2004, 131, 2329-2338; Mech Dev, 2004, 121, 1425-1441; Dev Biol, 2007; 307, 483-497). Известные мутации в этом гене у мыши и человека имеют рецессивный характер, но в гомозиготном состоянии приводят к серьезным аномалиям развития мозга, в диапазоне от недоразвития гипофиза и дисплазии оптического нерва и перегородки больших полушарий до слияния желудочков конечного мозга и отсутствия его структур. Поэтому исследователи предположили, что возникновение гена Anf у предков позвоночных могло послужить одной из ключевых предпосылок к возникновению конечного мозга в эволюции (Dev Biol, 2007; 307, 483-497). Поскольку Anf пока не был обнаружен у представителей самой древней группы современных позвоночных – у бесчелюстных (миноги и миксины), – одной из задач, решаемых в настоящее время в Лаборатории, является поиск этого гена у миноги.
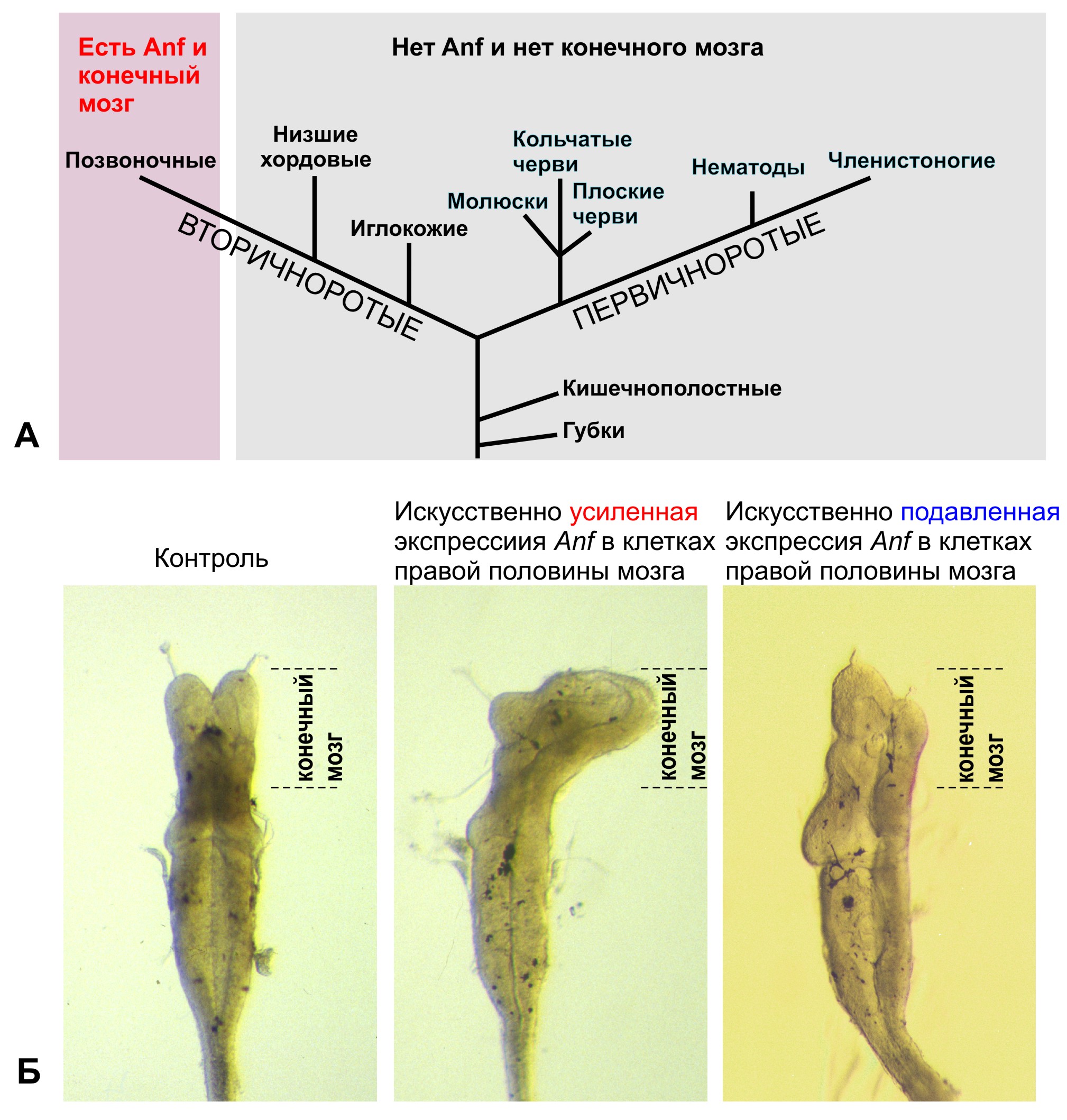
А. Гомеобоксный ген Anf и конечный мозг присутствуют только у позвоночных животных, в том числе у человека, и отсутствуют у всех остальных животных. Б. Искусственное изменение уровня экспрессии гомеобоксного гена Anf вызывает изменения размеров конечного мозга у головастика шпорцевой лягушки.
Лаборатория также изучает роль генов, пропавших в ходе эволюции позвоночных, – в регуляции регенерации. Исследователи установили, что некоторые из найденных генов-мишеней гомеобоксного гена Anf (гены секретируемого фактора Ag1 и гены малых ГТФаз Ras-dva) присутствуют только в геномах низших позвоночных, рыб и амфибий, но отсутствуют у высших, у рептилий, птиц и млекопитающих (Gene Expr Patterns, 2003, 325-30; Development, 2006, 133, 485-494; Nucleic Acids Res, 2006, 34, 2247-2257; Gene Expr Patterns, 2011, 11, 156–161). При этом у низших позвоночных эти гены, кодирующие разные типы белков, регулируют два важнейших процесса – развитие мозга и регенерацию конечностей (Sci Rep, 2013, 3, 1279; Biol Open, 2014, 3, 192-203; Sci Rep, 2015, 5:8123). Это дало основание выдвинуть гипотезу о том, что исчезновение в эволюции найденных генов могло быть своеобразной платой, сделанной предками высших позвоночных за возможность прогрессивного развития мозга. В рамках этого проекта сотрудники с коллегами разработали алгоритм и программу, которые позволяют на основе сравнения геномов идентифицировать гены, возникшие или исчезнувшие на заданном этапе эволюции. Функции некоторых из генов, найденных с помощью этой программы, сейчас изучаются в Лаборатории.

Инъекции анти-смыслового морфолинового олигонуклеотида блокируют трансляцию мРНК гена Ag1, что приводит к ингибированию регенерации
Изучение генной сети, связанной с функционированием гена Anf в клетках раннего зачатка переднего мозга, привело к открытию ряда неизвестных ранее генов, играющих важную роль в эмбриогенезе. Так, были открыты и изучены регуляторы раннего развития головного мозга – секретируемые белки Noggin2 и Noggin4 (Gene Expr Patterns, 2006 6:180-6; Development, 2011, 138, 5345-5356; Int J Dev Biol, 2012;56: 403-6; Sci Rep, 1996, 14, 6: 23049). Эти белки в отличие от их широко известного гомолога, ингибитора BMP – белка Noggin1, имеют способность связывать и ингибировать функцию регулятора задних отделов центральной нервной системы – секретируемого белка Wnt8.
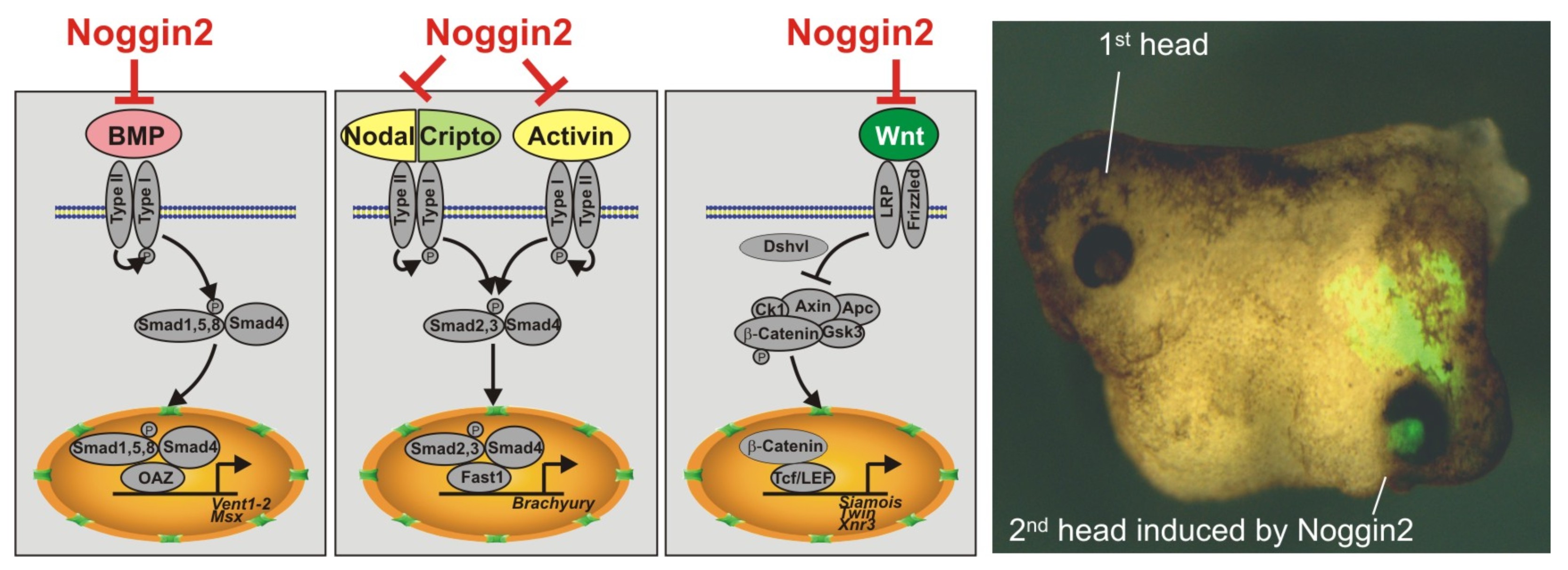
Секретируемый белок Noggin2 ингибирует три сигнальных каскада - BMP, Activin/Nodal и Wnt, что является достаточным условием для индукции второй головы при искусственно вызванной экспрессии Noggin2 на брюшной стороне эмбриона шпорцевой лягушки.
Кроме того, исследователи разработали ряд методов для изучения диффузии и взаимодействий секретируемых белков-морфогенов в межклеточном пространстве эмбриональных тканей (Sci Rep, 2016, 6:23049; Biochem Biophys Res Commun, 2015; 468:331-6). Впервые измерили коэффициенты диффузии белков семейств Noggin и Wnt in vivo и показали роль адсорбции на внеклеточном матриксе в их диффузии. Впервые с помощью математического моделирования показали, что такая адсорбция может играть роль важного фактора, необходимого для создания пространственно-упорядоченных структур в эмбриогенезе.
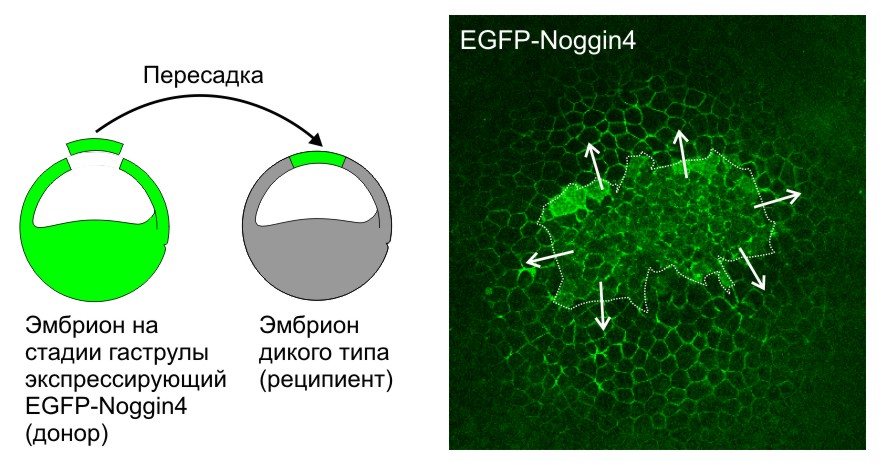
Секретируемый белок Noggin4, меченный EGFP (EGFP-Noggin4), диффундирует из пересаженного кусочка эктодермы (граница выделена пунктиром), клетки которого экспрессируют EGFP-Noggin4, по межклетникам эмбриона-реципиента на большие расстояния. Стрелки показывают направление диффузии.
Также в Лаборатории впервые была изучена роль в раннем эмбриональном развитии цитоскелетного белка Zyxin, который способен связываться с гомеодоменным белком Anf и с эффектором Shh сигнального каскада – транскрипционным фактором Gli1, ингибируя его активность (Dev Dyn, 2008, 237, 736-749; Dev Biol, 2013, 380, 37-48). Полученные данные важны, так как они впервые демонстрируют связь между одним из цитоскелетных белков-регуляторов морфогенетических движений клеток и важным сигнальным каскадом, управляющим дифференцировкой клеток в нейральном зачатке.
В Лаборатории проводятся работы по тестированию возможности использования генетически кодируемых флуоресцентных репортеров и сенсоров для изучения различных процессов в раннем эмбриогенезе и при регенерации (Nat Biotechnol, 1999, 17, 969-973; Science, 2000, 290:1478-1479; Nat Biotechnol, 2003, 21:191-194; Nat Methods, 2007, 4, 741-746; Nat Methods, 2010, 7, 827-829; Nat Commun, 2012 13; 3 :1204). В частности, впервые была показана возможность использования флуоресцентных белков из коралловых полипов для прижизненного мониторинга клеток в эмбрионах шпорцевой лягушки (Nat Biotechnol, 1999, 17, 969-973).
Аквариальная комната, где содержатся лягушки и головастики некоторых трансгенных линий, а также взрослые лягушки, экспрессирующие флуоресцентные белки под контролем промоторов различных генов.
Лаборатория сотрудничает с другими лабораториями Института, а также с кафедрой биофизики МГУ, кафедрой эмбриологии МГУ, Центром «Биоинженерия» РАН, НИИ физико-химической биологии им. А.Н. Белозерского МГУ, Институтом проблем передачи информации РАН, НИЦ Курчатовский институт, Массачусетским технологическим институтом (США), Университетом Виргинии (США).
Лаборатория была образована в 2005 году на основе группы с аналогичным названием, выделившейся в 1995 году из Лаборатории структуры и функции генов человека.
- Изучение молекулярно-генетических механизмов раннего развития и эволюции нервной системы.
- Изучение процессов регенерации больших придатков тела на моделях низших позвоночных.
- Проведение уникальных для нашей страны работ по созданию и поддержанию линий трансгенных лягушек Xenopus, экспрессирующих генетически кодируемые флуоресцентные репортеры и сенсоры.
- Установлено, что некоторые из найденных генов-мишеней гомеобоксного гена Anf (гены секретируемого фактора Ag1 и гены малых ГТФаз Ras-dva) присутствуют только в геномах низших позвоночных, рыб и амфибий, но отсутствуют у высших, у рептилий, птиц и млекопитающих.
- Открыты и изучены регуляторы раннего развития головного мозга – секретируемые белки Noggin2 и Noggin4.
- Разработан ряд методов для изучения диффузии и взаимодействий секретируемых белков-морфогенов в межклеточном пространстве эмбриональных тканей.
- Впервые была изучена роль в раннем эмбриональном развитии цитоскелетного белка Zyxin, который способен связываться с гомеодоменным белком Anf и с эффектором Shh сигнального каскада – транскрипционным фактором Gli1, ингибируя его активность.
- Разработана технология двухцветного репортерного вектора, существенно повышающая эффективность функционального промоторного анализа, благодаря возможности сравнивать экспрессионный потенциал двух делеционных мутантов изучаемого генного промотора в одном и том же трансгенном эмбрионе (Development, 2004, 131, 2329-2338).
- С помощью генетически кодируемого pH-индикатора SypHer2 на модели головастиков шпорцевой лягушки обнаружен ранее неизвестный эффект – быстрое закисление цитоплазмы клеток вблизи места ампутации хвоста (Biochim Biophys Acta, 2015 1850: 2318-28). Этот эффект является одной из первых реакций организма на ампутацию и может иметь важное регуляторное значение для последующей регенерации.
| ФИО | Должность | Контакты |
|---|---|---|
| Зарайский Андрей Георгиевич, проф., д.б.н. | г.н.с. | |
| Байрамов Андрей Вячеславович, д.б.н. | в.н.с. | |
| Ерошкин Федор Михайлович, к.б.н. | с.н.с. | xenopus.fe@gmail.com |
| Мартынова Наталья Юрьевна, к.б.н. | с.н.с. | martnat61@gmail.com, +7(916)1811632 |
| Ермакова Галина Владимировна | н.с. | |
| Орлов Евгений Евгеньевич, к.б.н. | н.с. | |
| Сетдекова (Паршина) Елена Анатольевна, к.б.н. | н.с. | |
| Терёшина Мария Борисовна, к.б.н. | н.с. | |
| Тимошина П.С. | м.н.с. | |
| Банникова М.А. | инж.-иссл. | |
| Волков С.А. | инж.-иссл. | |
| Колюпанова Н.М. | тех.-лаб. | |
| Курдюмов А.М. | тех.-лаб. | |
| Ватанина Н.И. | инженер | |
| Серебрякова М.В. | инженер | |
| Фирсова Т.В. | инженер | |
| Шитиков А.Д. | м.н.с. | |
Ранее здесь работали | ||
| Иванова Анастасия Сергеевна, к.б.н. | ||
| Нестеренко Алексей Михайлович | ||
| Короткова Д.Д., к.б.н. | ||
| Бородулин Александр Владиславович, к.б.н. | ||
| Арасланова К.Р. | ||
| Иванова Э.Д. | ||
| Карпова Л.А. | ||
| Аверьянова О.В. | ||
| Соловьева Е.А. | ||
 Загрузка...
Загрузка...Научные проекты
 Загрузка...
Загрузка...Зарайский Андрей Георгиевич
Москва, ул. Миклухо-Маклая, 16/10 — На карте
Scopus: 7003375442, ORCID: 0000-0003-4681-8169
 Загрузка...
Загрузка...
