Лаборатория биомолекулярной ЯМР-спектроскопии

|
Руководитель: Бочаров Эдуард Валерьевич |
Лаборатория исследует структуру белков и пептидов. Для этого используют один из важнейших современных методов – спектроскопию ядерного магнитного резонанса (ЯМР).
Сотрудники лаборатории занимаются исследованиями мембранных белков, таких, как рецепторные тирозинкиназы, ионные каналы, toll-подобные рецепторы, предшественник бета-амилоида, GPCR-ы и другие. Ведутся работы по изучению свойств природных люцеферинов, механизмов действия блокаторов болевых рецепторов, вирусных белков, необходимых для заражения клетки, а также механизмов лиганд-рецепторных взаимодействий.
Большая часть исследований напрямую связана с практическими вопросами: поиском противораковых мишеней, причин возникновения болезни Альцгеймера, созданием эффективных болеутоляющих, специфичных диагностических систем и др.
В распоряжении лаборатории находятся самые современные приборы фирмы Bruker: 600, 700 и 800 MHz, оборудованные криодатчиками, имеется твердотельный датчик с вращением под «магическим» углом. Кроме того, в лаборатории есть необходимое оборудование и технологии для бактериального и бесклеточного синтеза рекомбинантных белков и их физико-химической характеризации. На их основе были разработаны методы получения изотопно-меченых и селективно-меченых белков.
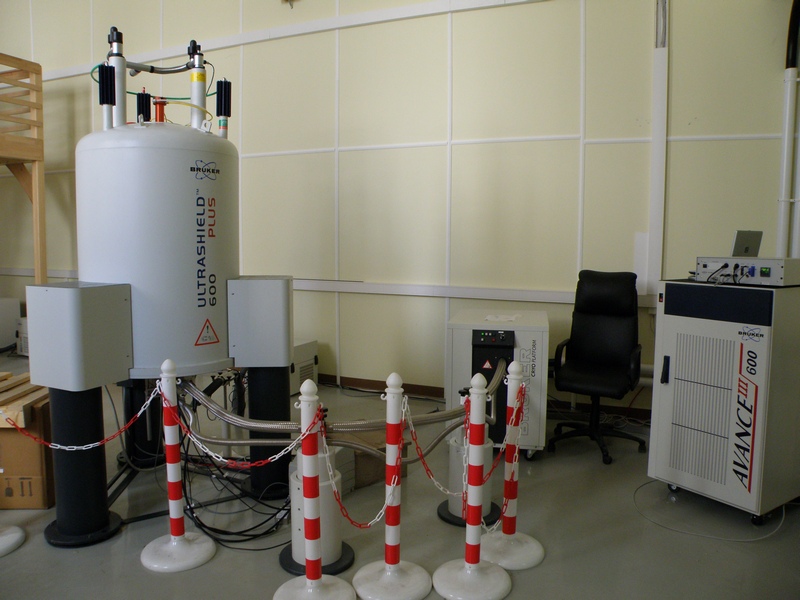
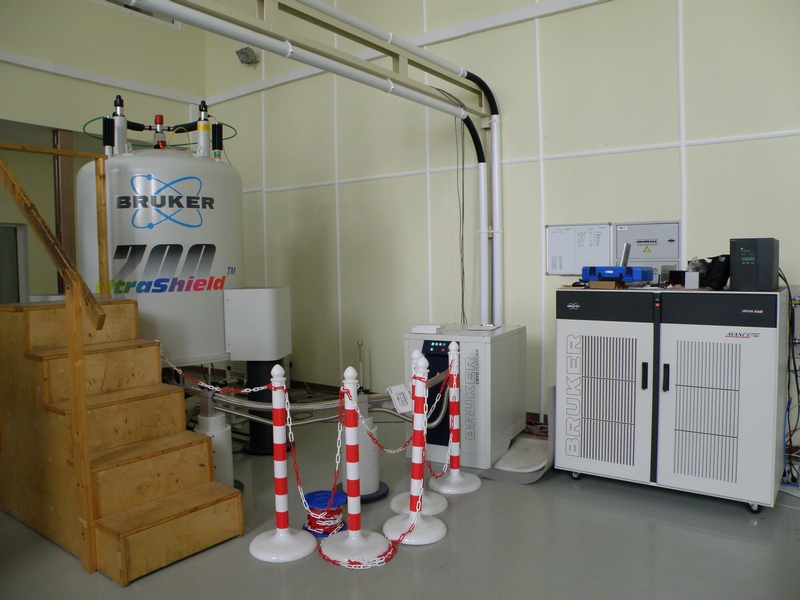
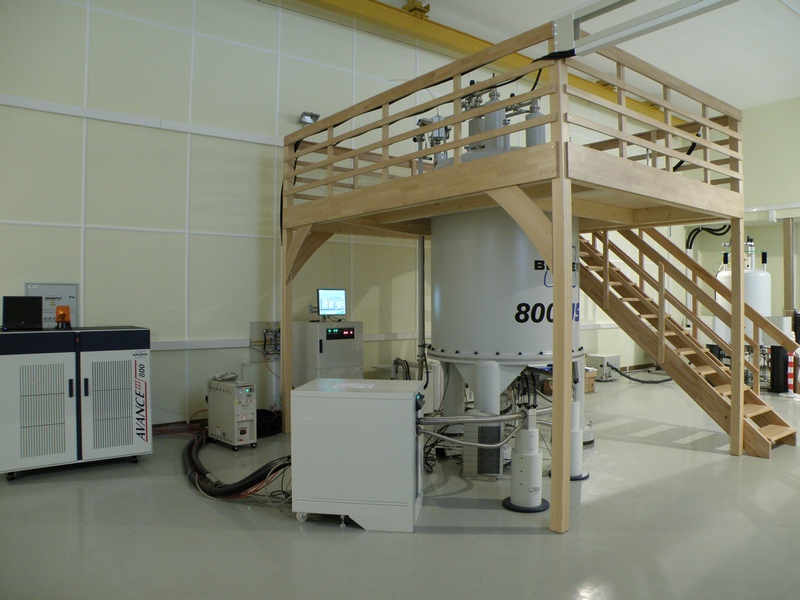
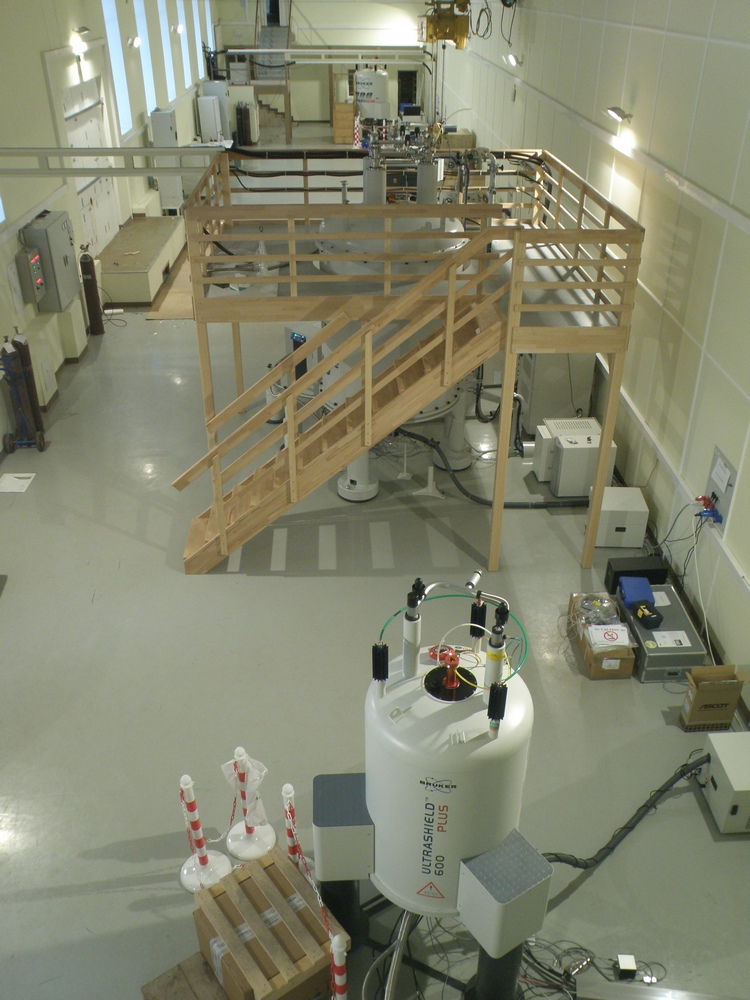




Все это позволяет успешно решать сложнейшие задачи на грани и за гранью возможностей современной структурной биологии.
Оборудование лаборатории является частью центра коллективного пользования ИБХ РАН, поэтому существует возможность выполнить анализ образцов высокой сложности методами ЯМР-спектроскопии на коммерческой основе.
Лаборатория имеет богатую историю. В 1965 году ее основал Владимир Федорович Быстров, член-корреспондент Академии наук СССР. Он одним из первых в мире начал заниматься структурными исследованиями белков и пептидов в растворе методом ЯМР-спектроскопии и создал крупнейшую в Советском Союзе научную школу. В 1990 г. лабораторию возглавил его ученик, профессор Александр Сергеевич Арсеньев.
По сей день лаборатория продолжает начинания Быстрова и является одной из ведущих мировых школ ЯМР в мире. Каждый год она выпускает новых высококвалифицированных специалистов и кандидатов наук, способных решать самые сложные задачи с использованием ЯМР-спектроскопии.
За долгие годы работы у лаборатории появилось множество друзей и партнеров. Среди них – лаборатория нобелевского лауреата Курта Вютриха, одна из сильнейших в мире ЯМР-лабораторий профессора Герхарда Вагнера, вторая по величине в мире фармацевтическая компания «Новартис» и другие.
Сегодня наша лаборатория – это очень дружный коллектив, который ставит перед собой самые амбициозные задачи и готов на любое интересное сотрудничество!
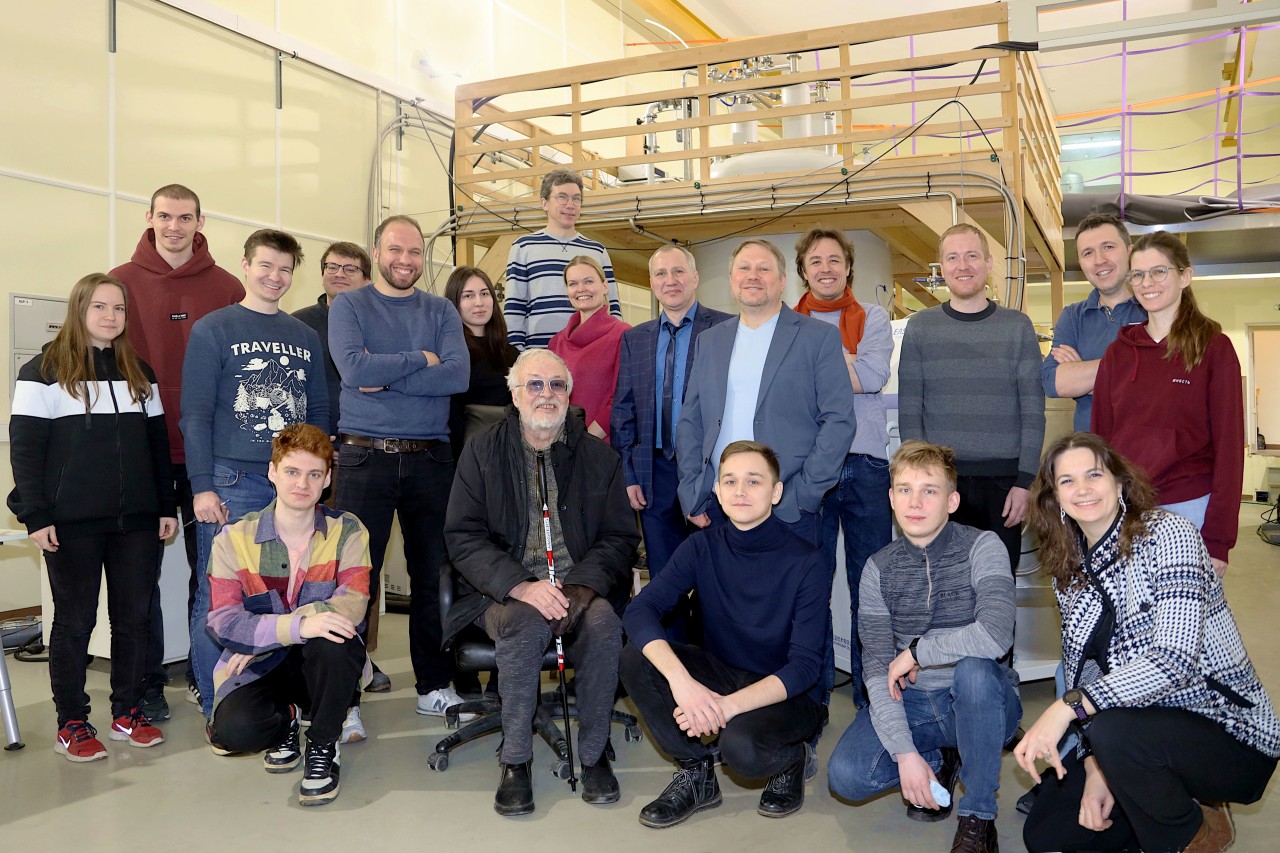
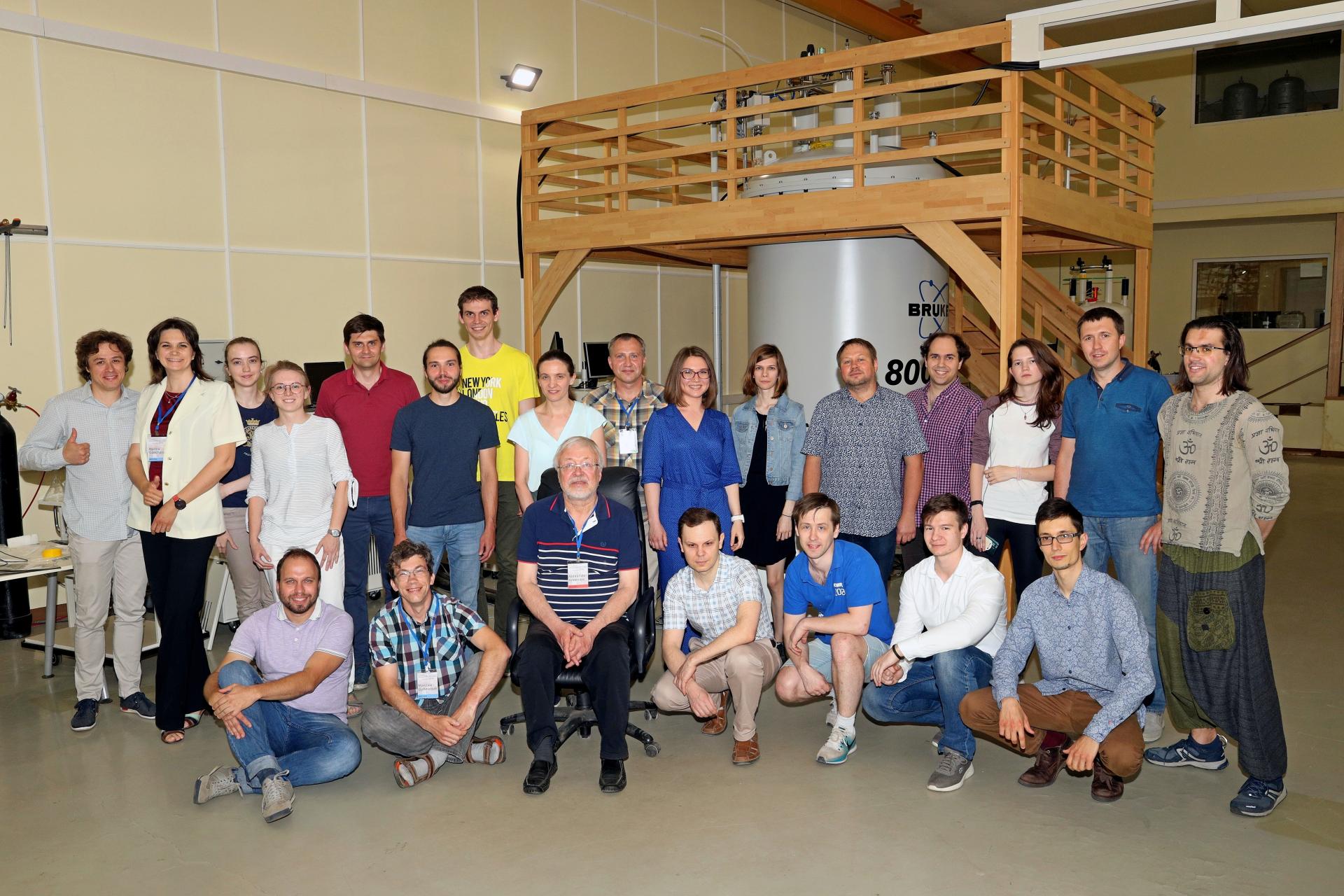
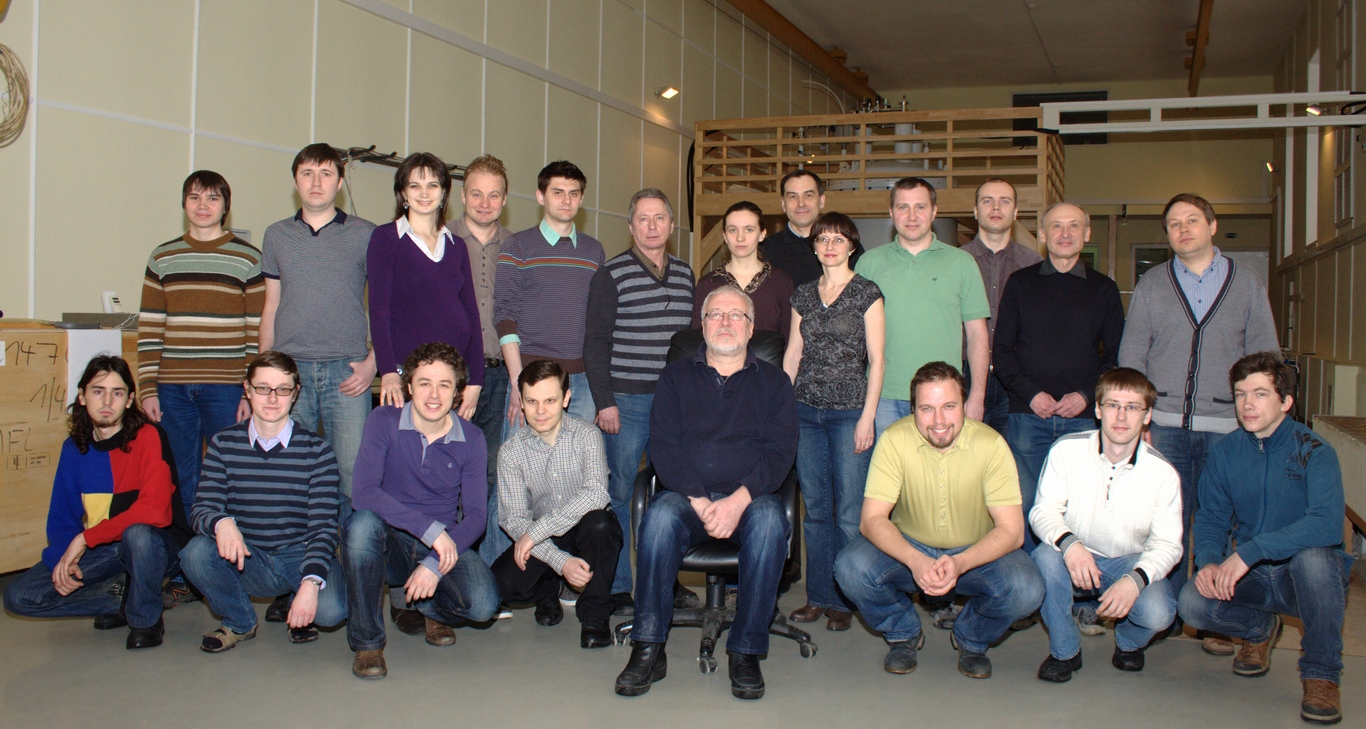
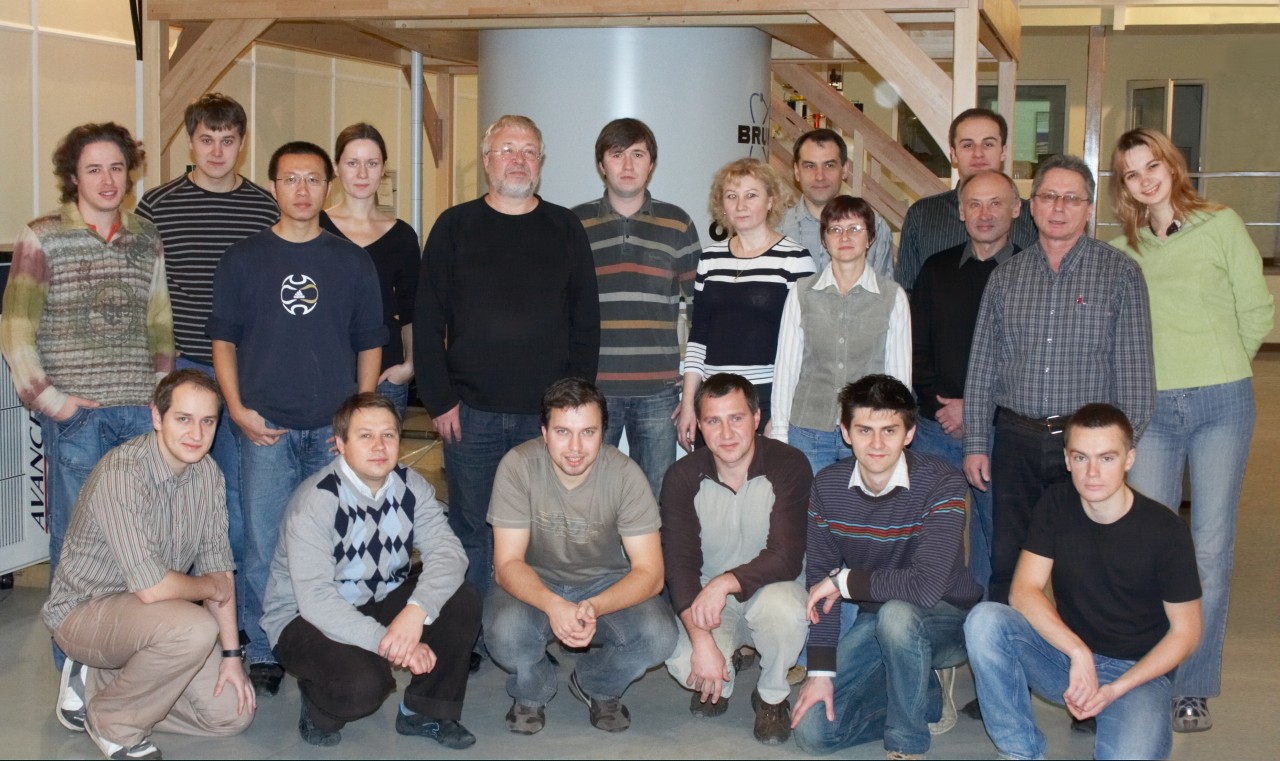




I. Биологические исследования
Установление структуры биомолекул в контексте присущих им взаимодействий:
- Пептиды защитной системы растений: гевеин-подобные и их взаимодействия с гликополимерами клеточной стенки грибов (хитином), липид-переносящий белок и его взаимодействие с фосфолипидами, ингибиторы трипсина и других протеаз
- Гомо- и гетеродимеразиция трансмембранных спиралей битопных мембранных белков (рецепторы тирозинкиназ)
- G-белок сопряжённые рецепторы
- Пептидные антибиотики и изучение механизма их действия
- Структура и межмолекулярные взаимодействия трансмембранных фрагментов "amyloid precursor protein" (APP), участвующего в развитии болезни Альцгеймера
- Мембраноактивные пептиды (токсины из яда змей, насекомых, фрагменты вирусных белков и др.) и их взаимодействия с биологическими мембранами.
II. Методические разработки для ЯМР- и ЭПР-спектроскопии биомолекул
- Разработка методов получения изотопно-меченых мембранных пептидов и белков
- Разработка новых мембрано-моделирующих сред для исследования мембранных белков;
- Программное обеспечение для анализа:
- Гетероядерных (13C,15N) ЯМР-данных белков и пептидов
- Диффузии макромолекул, формы линии 31Р-ЯМР (2Н-ЯМР) широких линий фосфолипидных дисперсий
- Декомпозиции двухкомпонентных спектров ЭПР.
Первым весомым вкладом в копилку мирового опыта исследования структуры пептидов и белков методом ЯМР стала работа коолектива под руководством В.Ф. Быстрова, устанавливающая взаимосвязь между константой спин-спинового взаимодействия протонов H-NCa-H и двугранным углом Q (Tetrahedron, 1973). Найденная зависимость получила название: уравнение Быстрова.
Исследования не ограничивались пептидами и белками. Прорыв был сделан и в мембранологии. Владимир Федорович совместно с лабораторией химии липидов ИБХ (руководитель – член-корреспондент АН СССР Бергельсон Л.Д.) нашёл способ измерения проницаемости фосфолипдных мембран с помощью дифференции их внешних и внутренних поверхностей с использованием парамагнитных ионов (Chemistry and Physics Lipids, 1971).
После отработки методологии последовательного отнесения сигналов в спектрах ЯМР, сконцентрированной в Швейцарии, в лаборатории проф. К. Вютриха, в которой принимал участие стажировавшийся там А.С. Арсеньев, несомненным успехом лаборатории Быстрова стали биологические приложения метода ЯМР.
В 1985 установили детальную пространственную структуру грамицидинового канала в комплексе с одновалентными катионами, выяснили механизм блокирования канала двухвалентными катионами, установили взаимосвязь дисперсности ионной проводимости и внутримолекулярной динамики, механизм открывания и закрывания канала. Работа, посвящённая структуре грамицидоновго канала (FEBS Letters, 1985), до сих пор является одной из самых цитируемых работ лаборатории.
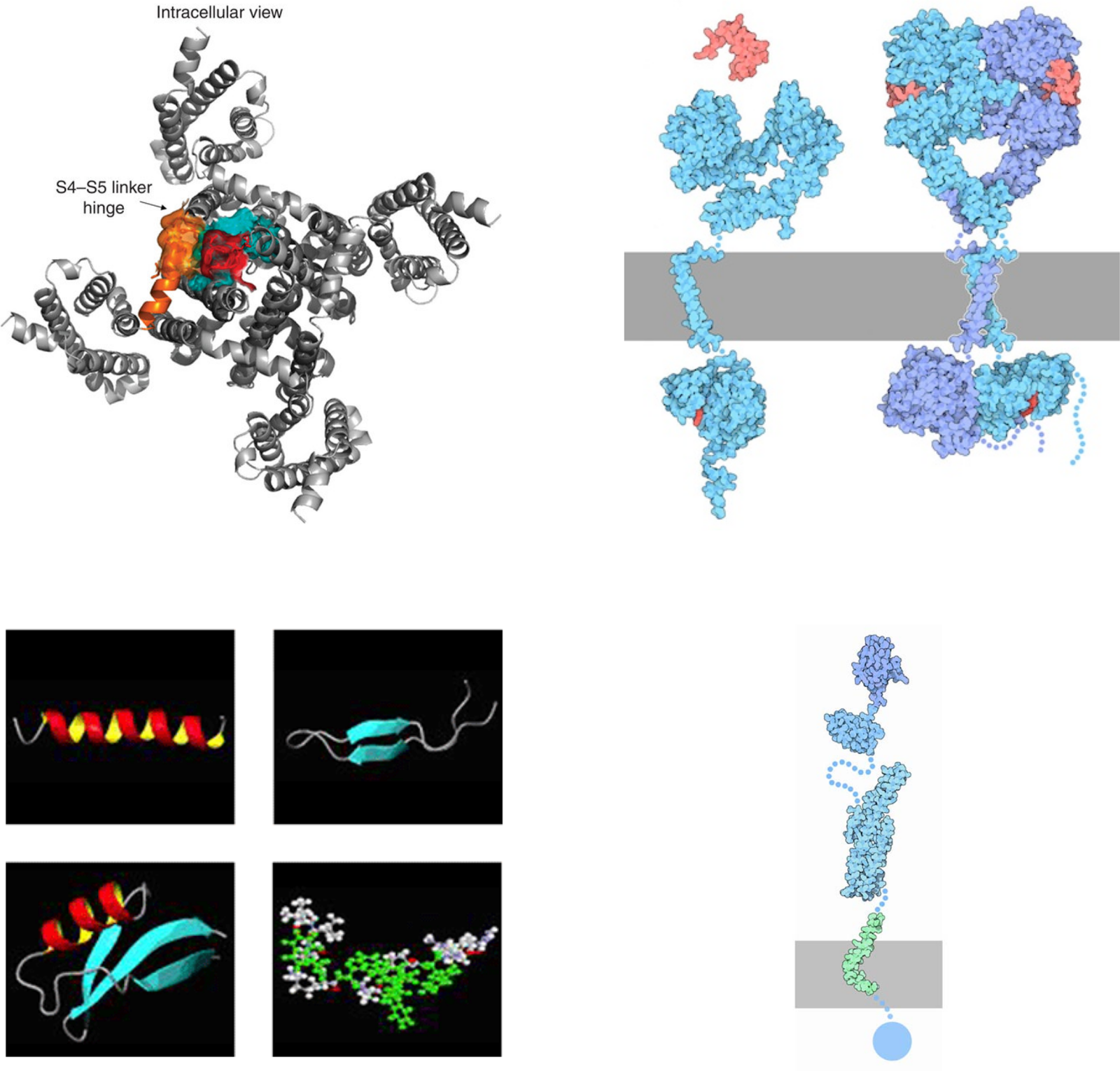
С тех пор "димерная" тема стала визитной карточкой Лаборатории. Как оказалось, многие биологические процессы сопровождаются димеризацией задействованных в них молекул полипептидов. В частности, именно димеризацией транс-мембранных доменов тирозинкиназных рецепторов сопровождается их активация (см. обзор, посвящённый этому направлению (Cell Adhesion and Migration, 2010)), нарушение димеризации транс-мембранного домена белка АРР отвечает за развитие болезни Альцгеймера человека (FEBS Letters, 2012), формирование транс-мембранных бета-шпилечных димеров ареницином лежит в основе антимикробной активности этого пептида (Biochemistry, 2011).
В настоящее время лаборатория располагает собственной базой для получения изотопно-меченых рекомбинантных белков генно-инженерными методами, современными ЯМР-спектрометрами и программным обеспечением для обработки и анализа получаемых спектров. Например, несомненным авторитетом среди ЯМР-спектроскопистов пользуется программа DASHA для анализа релаксационных данных биомолекул, разработанная под руководством проф. А.С. Арсеньева (Applied Magnetic Resonance. 1995). Постоянный приток молодых сотрудников в лабораторию и её популярность среди студентов МФТИ позволяют надеяться, что список достижений лаборатории будет пополняться.
 Загрузка...
Загрузка...Научные проекты
 Загрузка...
Загрузка...Бочаров Эдуард Валерьевич
Москва, ул. Миклухо-Маклая, 16/10 — На карте
ResearcherID: R-5231-2016, Scopus: 7004085574, ORCID: 0000-0002-3635-1609, Google Scholar
 Загрузка...
Загрузка...
