Лаборатория структурной биологии ионных каналов
|
Руководитель: Шенкарёв Захар Олегович |
На фото слева направо: Михаил Мышкин, Захар Шенкарёв, Андрей Царёв, Милита Кочаровская, Александр Парамонов.
Лаборатория исследует мембранные рецепторы и ионные каналы, а также соединения, действующие на них (токсины, эндогенные лиганды). Для исследования структуры и механизмов действия на молекулярном уровне мы применяем современные методы структурной биологии: ЯМР-спектроскопию и криоэлектронную микроскопию. В настоящее время мы изучаем механизмы взаимодействия токсинов с потенциал-зависимыми K+ и Na+ каналами (Kv и Nav), а также исследуем структуры белковых лигандов никотинового ацетилхолинового рецептора (nAChR). Структурные исследования мембранных рецепторов и каналов необходимы для разработки новых средств диагностики и терапии заболеваний нервной, сердечно-сосудистой и мышечной систем организма. Кроме того, мы исследуем структуры и механизмы действия антибиотиков и защитных пептидов растений и животных. Эти исследования необходимы для создания новых препаратов для лечения инфекционных заболеваний.
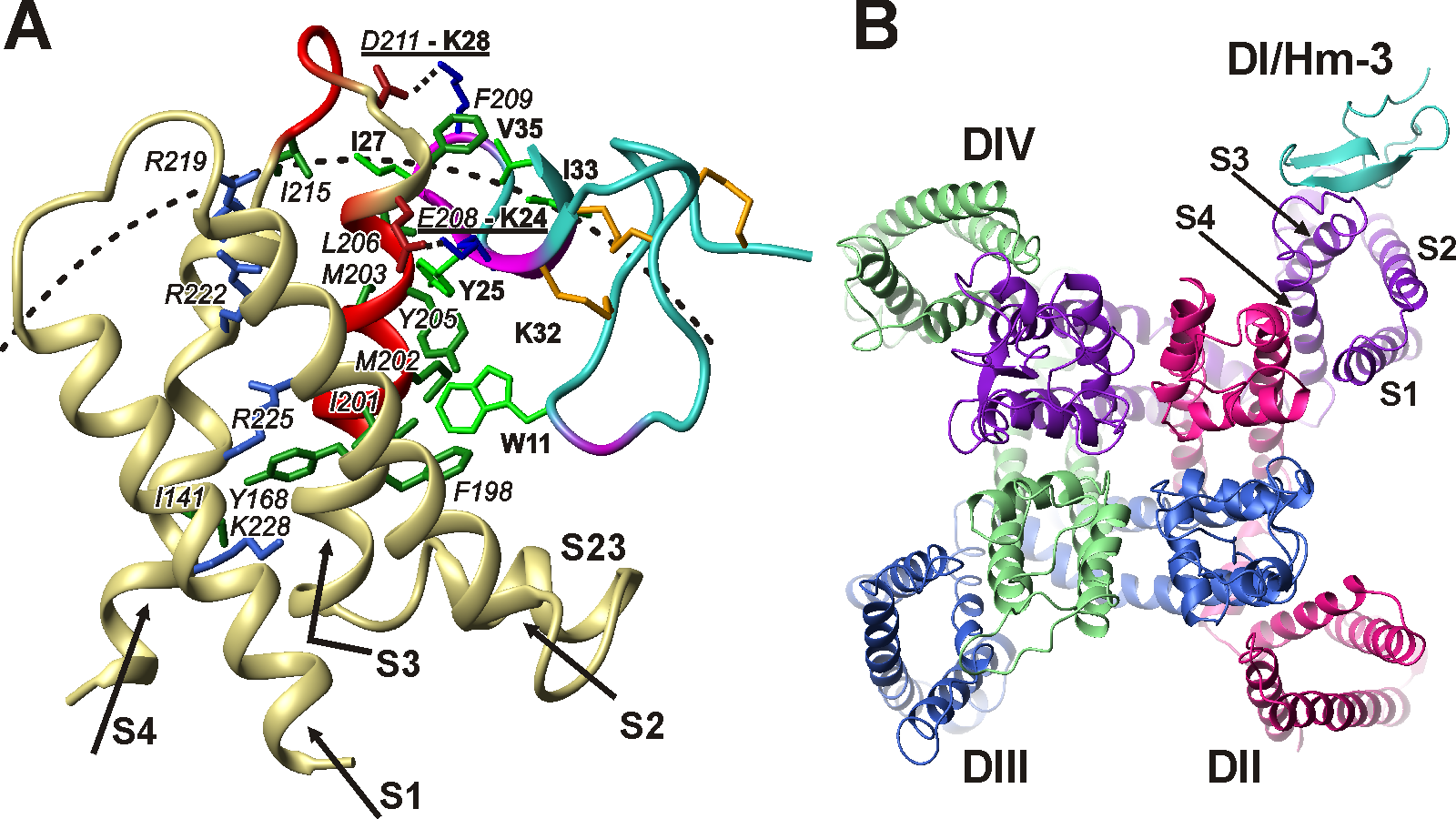
Рис 1. Структура комплекса канала Nav1.4 из мышц человека с токсином Hm-3 из яда паука Heriaeus melloteei по данным ЯМР. (A) Комплекс Hm-3 (голубой/фиолетовый) с первым потенциал-чувствительным доменом (DI) канала (песчаный/красный). Вид сбоку, со стороны липидного бислоя. (В) Комплекс токсин-канал. Вид на плоскость мембраны со стороны внеклеточного пространства.
Нашей лабораторией проведены ЯМР-исследования структуры и динамики потенциал-чувствительных доменов нескольких K+ и Na+ каналов, включая канал Nav1.4 из мышц человека. Дисфункции канала Nav1.4 вызывают расстройства опорно-двигательного аппарата, такие как паралич, миастенический синдром и миотония. Впервые охарактеризованы «необычные» флуктуации в структуре доменов, идущие с характерными временами в диапазоне мкс-мс, возможно, являющиеся прообразом структурных перестроек, происходящих при потенциал-зависимой активации. Показано, что домены Na+ каналов обладают большей конформационной пластичностью по сравнению с доменами K+ каналов. Это свойство, вероятно, обуславливает более быстрое срабатывание Nav каналов при изменении трансмембранного потенциала. Нами исследованы токсины VSTx1 и Hm-3 из яда пауков, которые, связываясь с участком мембраны, окружающий ионный канал, блокируют потенциал-зависимую активацию. На основании экспериментальных данных ЯМР-спектроскопии, совместно с лабораторией молекулярных инструментов для нейробиологии ИБХ РАН построена модель комплекса токсина Hm-3 с каналом Nav1.4 из мышц человека. Полученные структурные данные открывают возможность дальнейших фармакологических разработок.
В настоящее время, совместно с биологическим факультетом МГУ, используя методы электронной микроскопии, мы исследуем пространственную структуру полноразмерного канала Kv7.1 сердечной мышцы человека. Дисфункции этого канала приводят к развитию наследственной аритмии – синдрома удлиненного интервала QT.
Совместно с группой биоинженерии нейромодуляторов и нейрорецепторов ИБХ РАН, используя методы ЯМР-спектроскопии, нами определены структуры нескольких «трехпетельных» белков, действующих на ионный канал никотинового ацетилхолинового рецептора (nAChR). Определены пространственные структуры токсина WTX из яда кобры, регуляторных белков Lynx1 и Lypd6 из нервной системы человека и белков SLURP-1 и SLURP-2, продуцируемых клетками эпителия человека. Эти молекулы имеют перспективы для разработки новых лекарств, нацеленных на лечение нейродегенеративных заболеваний.
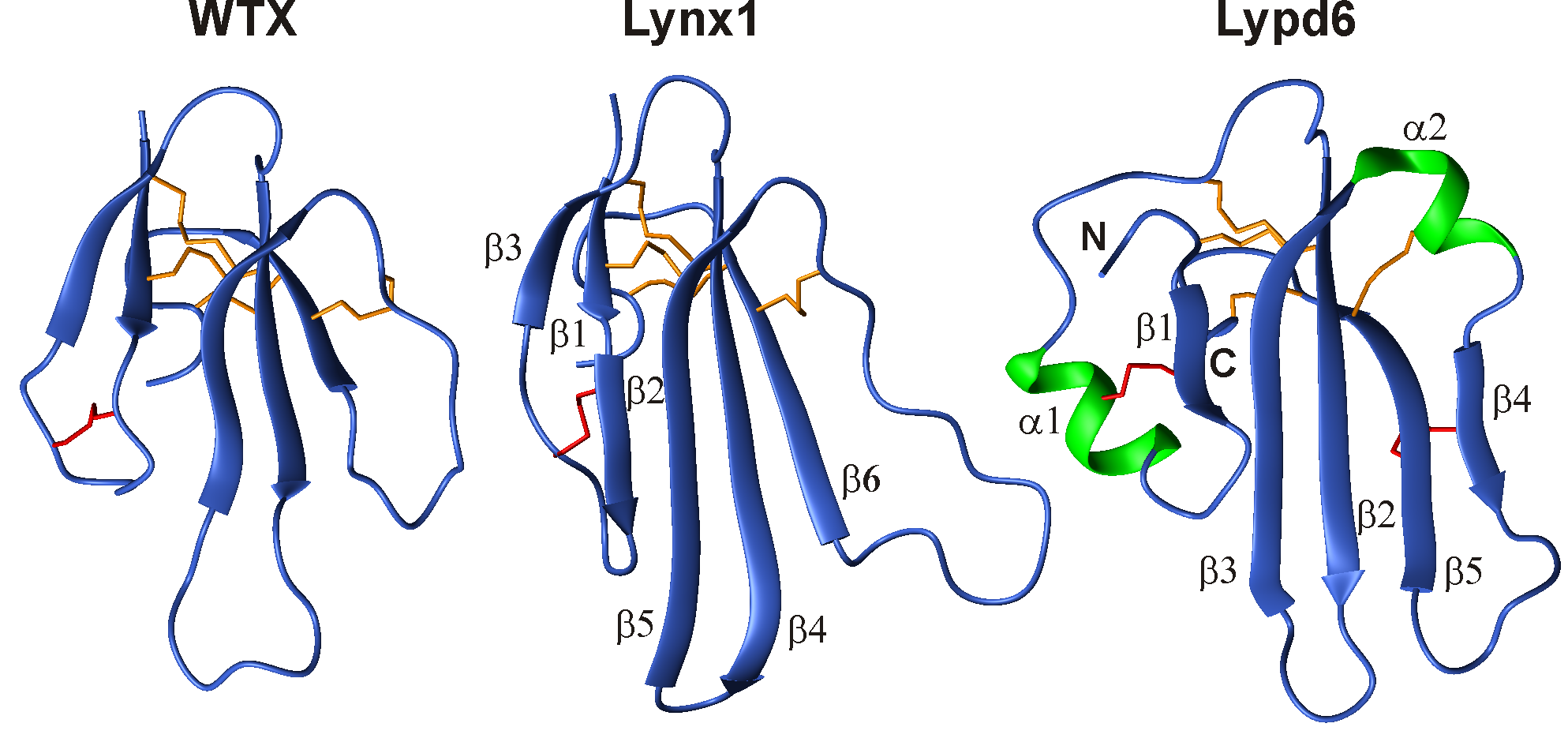
Рис 2. Пространственные структуры трехпетельных белков, действующих на никотиновые ацетилхолиновые рецепторы, по данным ЯМР. WTX токсин из яда кобры Naja kaouthia. Lynx1 и Lypd6 – регуляторные белки нервной системы человека.
Используя методы ЯМР-спектроскопии, наша лаборатория совместно с Учебно-научным центром ИБХ РАН исследует структуру и механизм действия новых антибиотиков и защитных пептидов растений и животных. Большинство из изученных молекул демонстрируют повышенное сродство к мембранам бактериальных клеток и способны образовывать поры и ионные каналы, которые и приводят к гибели клетки-мишени. Нами были определены пространственные структуры и исследована динамика следующих молекул: двухкомпонентный лантибиотик лихеницидин, антибиотик антиамебин, антимикробный пептид ареницин из морского червя, антимикробный пептид аурелин из медузы, пептид дефенсин из чечевицы, липид-транспортирующий белок чечевицы в комплексе с липидами.
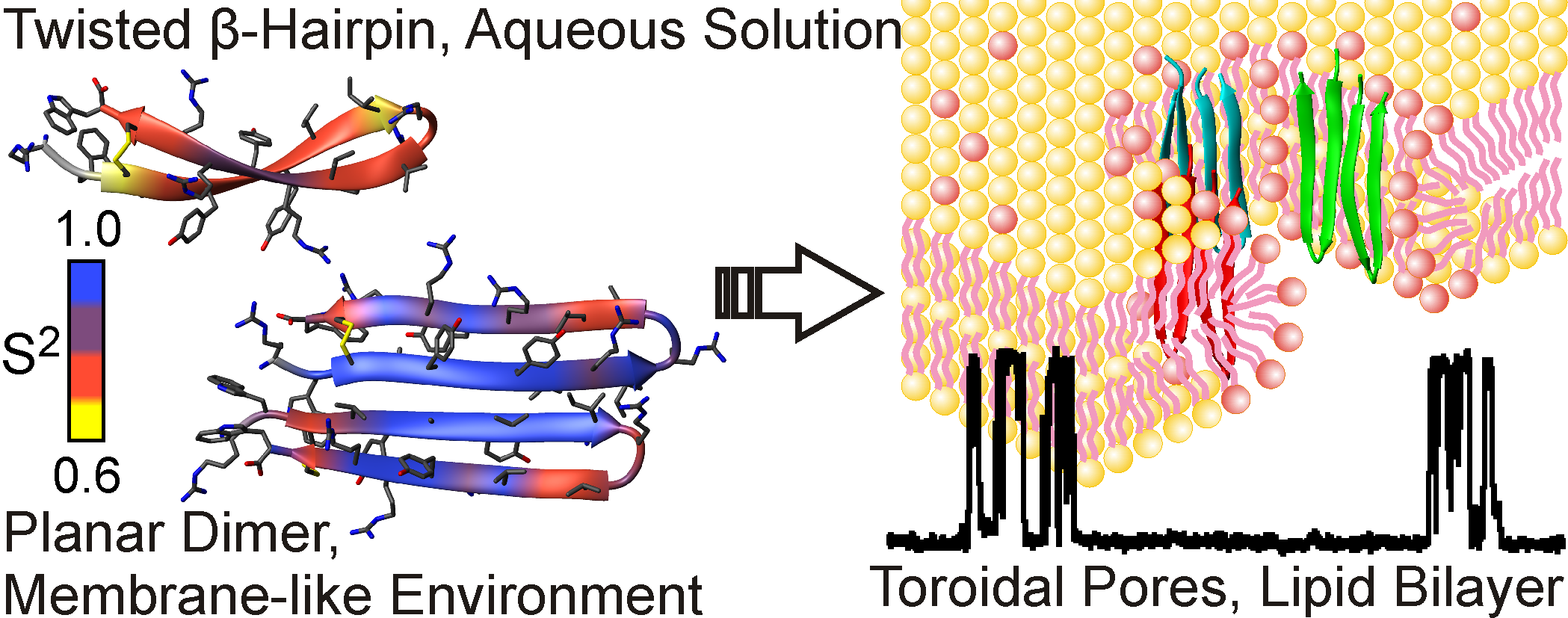
Рис 3. Пространственная структура и конформационная пластичность мономера антимикробного пептида ареницина в воде и димера пептида в мембраноподобном окружении. Справа показано образование ареницином тороидальных пор в мембране бактериальной клетки.
Также лаборатория разрабатывает новые подходы для изучения мембранных биомолекул. Одним из перспективных направлений является использование наночастиц на основе липопротеинов высокой плотности (нанодисков). Нами впервые была показана возможность применения нанодисков в качестве среды для ЯМР-исследований мембранных белков и мембраноактивных пептидов, исследована возможность применения нанодисков в бесклеточных системах синтеза для получения функционально-активных форм мембранных белков, а также возможность применения нанодисков для фолдинга мембранных белков in vitro.
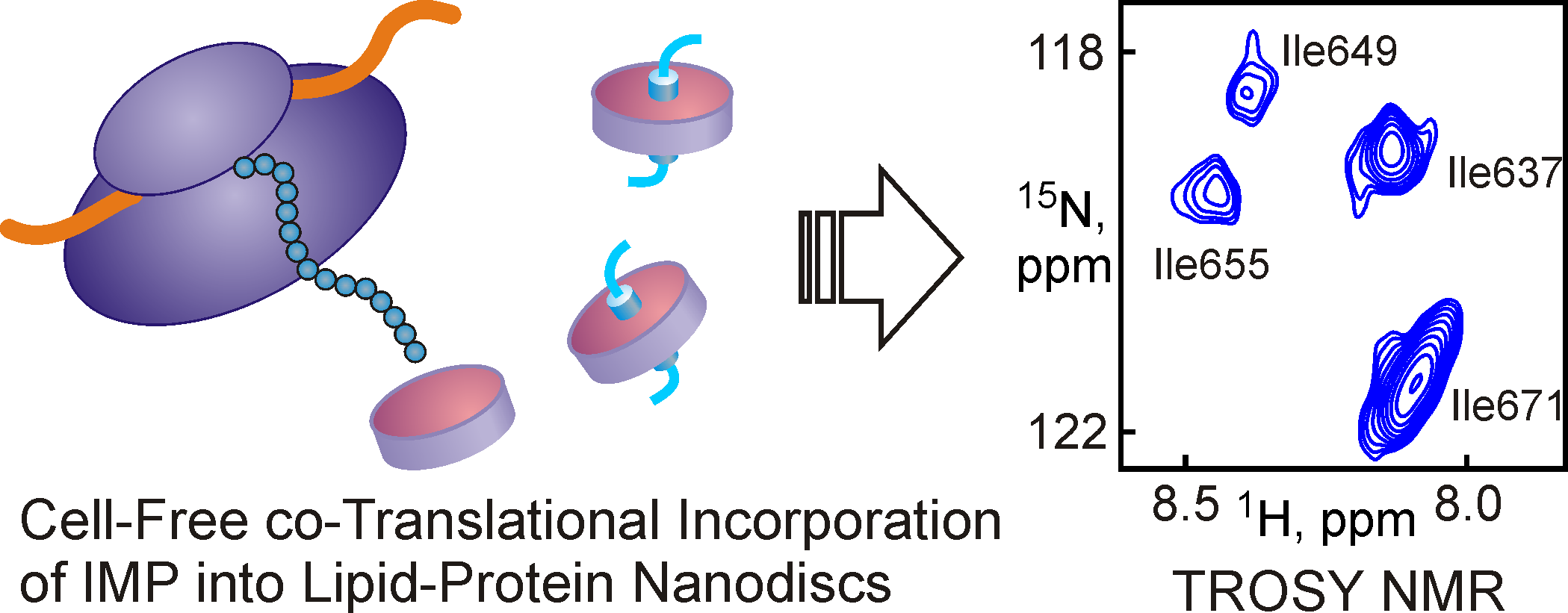
Рис 4. Добавление предварительно сформированных нанодисков в трансляционную смесь бесклеточной системы синтеза позволяет получать функционально-активные мембранные белки, доступные для исследования методами ЯМР-спектроскопии.
Нашей лабораторией ведутся разработки новых подходов для установления химической структуры и изучения механизмов превращения в азот-богатых гетероциклических соединениях. Один из разработанных подходов основан на измерении и анализе «малых» (амплитуда от 1 до 0.01 Гц) констант спин-спинового взаимодействия 13С-15N и 1H-15N в соединениях, селективно меченных стабильным изотопом 15N. Применение этого подхода позволило изучить процессы азидо-тетразольной таутомерии в азидо-триазинах и азидо-пиримидинах.
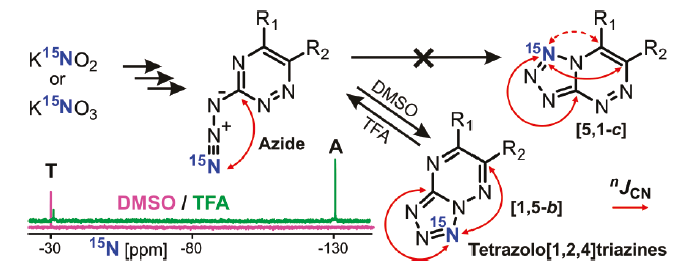
Рис 5. Анализ «малых» констант спин-спинового взаимодействия 13С-15N в соединениях, меченных изотопом 15N, позволяет однозначно установить структуру тетразольной формы в азидо-триазинах и азидо-пиримидинах.
Лаборатория ведет работы в сотрудничестве с подразделениями ИБХ РАН:
- Группа биоинженерии нейромодуляторов и нейрорецепторов (Руководитель – Люкманова Е.Н.)
- Лаборатория биомолекулярной ЯМР-спектроскопии (Руководитель – Арсеньев А.С.)
- Лаборатория молекулярных инструментов для нейробиологии (Руководитель – Василевский А.А.)
- Учебно-Научный центр ИБХ РАН (Руководитель – Овчинникова Т.В.)
- Лаборатория молекулярного дизайна и синтеза ИБХ РАН (Руководитель – Коршун В.А.)
Лаборатория сотрудничает с Российскими научными и образовательными организациями:
- Кафедра Биоинженерии биологического факультета МГУ (группа структурной биотехнологии, руководитель проф. Соколова О.С.)
- Лаборатория магнитного резонанса, Институт «Международный томографический центр» СО РАН, Новосибирск
- Кафедра органической и биомолекулярной химии Уральского федерального университета
Международные партнеры:
- Лаборатория профессора Александра Соболевского, Колумбийский университет (США). Sobolevsky Lab, Department of Biochemistry and Molecular Biophysics, Columbia University (USA). Head, Associate Professor Alexander Sobolevsky
Группа структурной биологии ионных каналов была организована в 2015 году в рамках программы Президиума РАН «Молекулярная и клеточная биология» (де факто группа существовала с 2008 года). В 2019 году группа получила статус лаборатории.
- Исследование структуры K+ и Na+ каналов человека и механизмов действия токсинов, влияющих на потенциал-зависимую активацию.
- Изучение взаимосвязи структура-функция в «трехпетельных» белках – лигандах никотинового ацетилхолинового рецептора.
- Исследование структуры и механизмов действия новых антибиотиков и защитных пептидов.
- Разработка новых мембраномоделирующих сред на основе липопротеинов высокой плотности (нанодиски) для продукции и структурно-функциональных исследований мембранных белков и пептидов.
- Разработка новых методов ЯМР для установления структуры и механизмов превращения азот-богатых гетероциклических соединений.
- Исследована структура и динамика двух потенциал-чувствительных доменов канала Nav1.4 из мышц человека. Исследована структура и мембранная активность токсина Hm-3 из яда паука. На основании экспериментальных данных ЯМР-спектроскопии построена модель комплекса Nav1.4/Hm-3.
- В потенциал-чувствительном домене канала KvAP археи охарактеризованы межспиральные движения в мкс-мс диапазоне, возможно, являющиеся прообразом структурных перестроек, происходящих при потенциал-зависимой активации.
- Определены пространственные структуры и исследована внутримолекулярная динамика для ряда «трехпетельных» белков, действующих на никотиновые ацетилхолиновые рецепторы, включая токсин кобры WTX, регуляторные белки Lynx1 и Lypd6 из нервной системы человека и белки SLURP-1 и SLURP-2 из эпителия человека.
- Определены пространственные структуры и предложен механизм действия для новых антибиотиков и защитных пептидов растений и животных, включая двухкомпонентный лантибиотик Лихеницидин; антибиотик антиамебин; антимикробные пептиды ареницин и аурелин, дефенсин и липид-транспортирующий белок чечевицы.
- Разработан ряд новых подходов по применению нанодисков для продукции и структурно-функциональных исследований мембранных белков и пептидов.
- Разработан новый метод ЯМР для установления химической структуры и изучения механизмов превращения в азот-богатых гетероциклических соединениях.
| ФИО | Должность | Контакты |
|---|---|---|
| Шенкарёв Захар Олегович, проф.РАН, д.ф.-м.н. | г.н.с. | zakhar-shenkarev@yandex.ru |
| Парамонов Александр Сергеевич, к.ф.-м.н. | с.н.с. | |
| Заиграев Максим Михайлович | инж.-иссл. | maximzaigraev@gmail.com |
| Горелов Станислав Александрович | инж.-иссл. | |
| Живов (Коваленко) Е.А. | инж.-иссл. | |
| Кочаровская Милита Владимировна | м.н.с. | |
| Миронов Павел Андреевич | м.н.с. | |
| Иванников А.Д. | инж.-иссл. | |
Ранее здесь работали | ||
| Лейченко Е.В. | ||
| Шабельников С.В. | ||
| Заняткин И.А., к.б.н. | ||
| Карлова М.Г. | ||
| Кветкина А.Н. | ||
| Мариловцева Е.В. | ||
| Мышкин Михаил Юрьевич | ||
| Попова А.С., к.б.н. | ||
| Царев А.В. | ||
| Хорев А.С. | ||
| Черная Елизавета Михайловна | ||
| Черников А.М. | ||
| Локтюшов Е.В. | ||
| Суркова Е.А. | ||
| Литвинов Д.С. | ||
| Орешков Сергей Денисович | ||
| Резникова О.В. | ||
| Уланова П.В. | ||
 Загрузка...
Загрузка...Научные проекты
 Загрузка...
Загрузка...Шенкарёв Захар Олегович
Москва, ул. Миклухо-Маклая, 16/10 — На карте
Scopus: 7801695075, Google Scholar, ORCID: 0000-0003-1383-3522
 Загрузка...
Загрузка...
