Пресс-центр / новости / Наука /
Структура супрамеров, сформированных амфифильными молекулами biot-CMG-DOPE
Ученые из Отдела химической биологии гликанов и липидов, Отдела структурной биологии и Отдела биоматериалов и бионанотехнологий ИБХ РАН совместно с российскими и зарубежными коллегами опубликовали в журнале ChemistryOpen работу, посвященную изучению супрамолекулярной организации конъюгата биотина с DOPE (диолеоилфосфатидилэтанламин) в составе клеточной мембраны и объяснению механизма его взаимодействия со (стрепт)авидином.
Молекула, формула которой показана на рисунке 1, представляет собой биотин, соединенный с DOPE (диолеоилфосфатидилэтанламин) нетривиальным спейсером, CMG(сделан из олигоглицина, часть азотов которого карбоксиметилирована). Этот спейсер мы придумали для того, чтобы делать водорастворимыми липофильные (из-за DOPE) молекулы, в том числе конъюгаты DOPE с гидрофобными пептидами. И эти самые конъюгаты, и biot-CMG-DOPE, замечательно встраиваются в живые клетки; а также за секунды, путем простого контакта, покрывают почти любые поверхности, - и это стало универсальным способом биотинилирования.
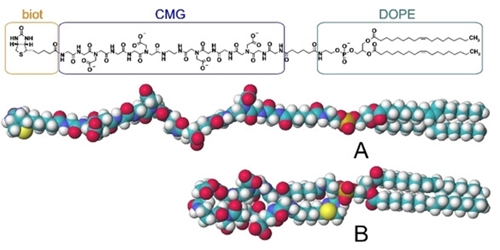
Рис.1. Структура молекулы biot‐CMG‐DOPE
Результаты молекулярной динамики (МД) говорят о том, что для biot-CMG-DOPE выгодна сложенная конформация В, а вовсе не распрямленная А. Но в такой конформации биотин должен быть совсем не «виден» для стрептавидина – ни в составе клеточной мембраны, ни в монослое, когда биотинилируется неживая поверхность. Однако, как уже сказано выше, biot-CMG-DOPE позволяет биотинилировать почти всё, и при этом биотиновый остаток прекрасно (всегда) взаимодействует со(стрепт)авидином. Как разрешить это противоречие? Для этого супрамеры, образуемые biot-CMG-DOPE, были изучены как экспериментально, так и с помощью МД. «Супрамеры» - это (здесь) глобулы, образуемые в водных средах, а также моно- или бислои на поверхностях. Физические методы дали согласованные результаты о размерах глобул, а метод малоуглового рентгеновского рассеяния (МУРР) позволил расшифровать организацию глобулы, показанную на рисунке 2.
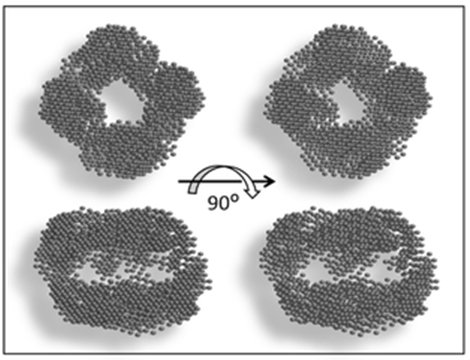

Рис.2. Организация biot‐CMG‐DOPE глобул
Кажущаяся полость – это ядро, в основном состоящее из DOPE, то есть, вполне ожидаемо глобула похожа на мицеллу. А вот понять, где расположены остатки биотина, помогли расчеты МД, см. рисунок 3.
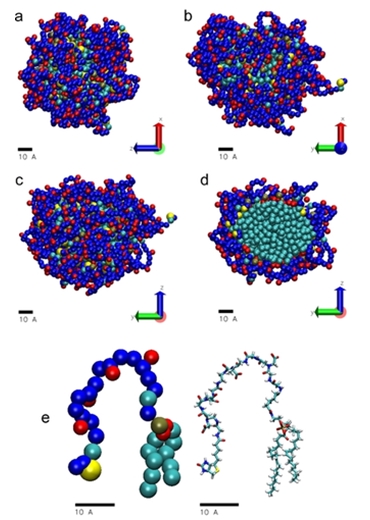
Рис.3. Результаты молекулярной динамики. Супрамеры biot-CMG-DOPE показаны в проекциях на плоскости XZ(a), XY (b) иYZ (d) и как поперечное сечение (с); (e) усредненная конформация отдельной молекулы: слева крупнозернистая модель, справа полноатомная модель. Показаны остатки биотина (желтый), фрагменты CMG (синий, СОО- группы, красный), фосфатные группы (коричневый), гидрофобные фрагменты (в основном остатки DOPE)(светло-голубой).
Согласно данным МД, большая часть биотиновых остатков тяготеет к гидрофобному ядру (конформация В на самом первом рисунке), но некоторое небольшое их количество оказывается на периферии и поэтому доступно для взаимодействия.
Но самое интересное для нас было понять устройство плоского слоя, сформированного biot-CMG-DOPE. Его организация в общих чертах похожа на расположение биотина, DOPE и CMG в глобуле, как показано на рисунке 4.
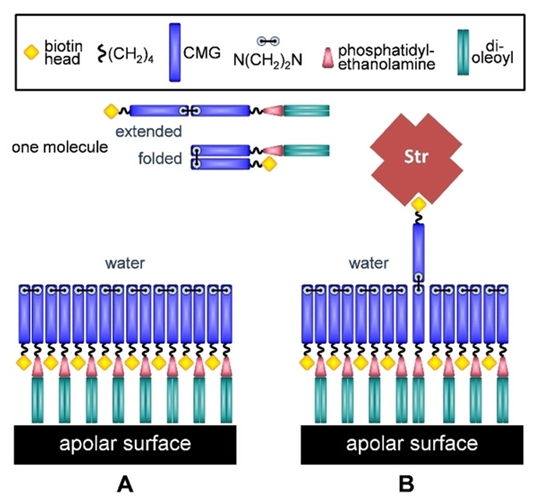
Рис.4. Схематическое представление плоского слоя, сформированного biot-CMG-DOPE (A - монослой на неполярных поверхностях; В - всплытие несвернутой молекулы и её связывание со (стрепт)авидином)
Однако здесь спрятанными оказываются практически все остатки биотина, и лишь каждый сотый способен динамически всплывать на поверхность слоя. Этого вполне достаточно, чтобы затем (и эксперимент это подтверждает) почти вся поверхность была покрыта монослоем (стрепт)авидина.
Почему всё это интересно?
biot-CMG-DOPE служит универсальным реагентом для быстрой и деликатной модификации живых клеток биотином и различными (стрепт)авидинилированными молекулами в физиологических условиях. Например, эритроциты, биотинилированные с помощью biot-CMG-DOPE, были живыми извлечены "за биотин" из крови после того, как они находились в кровотоке в течение 24 часов, что открывает горизонты для терапевтических технологий in vivo, основанных на модификации клеток biot-CMG-DOPE. Поэтому важно понять супрамолекулярную организацию этой молекулы в составе клеточной мембраны, а также ее потенциальную способность «скрывать» биотиновые остатки от биотин-разрушающего фермента крови (а такой есть) и биотин-специфических природных антител (которые тоже известны), в то же время оставляя возможность для взаимодействия.
6 июня 2020 года



