Пресс-центр / новости / Наука /
WINIGRET: метод широкомасштабной идентификации новых и потерянных генов, ответственных за эволюционные преобразования
Сотрудники Лаборатории молекулярных основ эмбриогенеза ИБХ РАН совместно с сотрудниками Института проблем передачи информации им. А.А. Харкевича разработали метод широкоформатной идентификации генов, потеря/возникновение которых в ходе эволюции была связана с потерей/появлением интересующего фенотипического или физиологического признака.
Большинство эволюционных преобразований в виде появления/исчезновения некоторого фенотипического или физиологического признака на том или ином этапе эволюции связаны либо с перестройками цис-регуляторных элементов, контролирующих генную активность, либо с изменениями непосредственно в белок-кодирующих последовательностях генов. Однако, в последнее время накапливается все больше данных, что значительная эволюционные изменения также могут быть связаны с потерей/возникновением целых генов. Целенаправленная широкоформатная идентификация таких генов оставалась из-за своей сложности до сих пор не решенной задачей. Между тем, выявление таких генов и дальнейшее изучение их функций важно не только для расшифровки фундаментальных основ эволюционных преобразований, но и для понимания механизмов возникновения множества генетических заболеваний.
Для решения этой задачи авторы данной статьи разработали метод WINEGRET, названный так, во-первых, по причине его образования из двух разнородных методов, а, во-вторых, по первым буквам краткого описания решаемой с помощью этого метода задачи: Wide-scale Identifcation of Novel/Eliminated Genes Responsible for Evolutionary Transformations. Главная идея метода заключается в том, чтобы искать нужные гены как гены удовлетворяющие одновременно двум требованиям: (1) они должны быть потеряны/появиться на том же этапе эволюции, на котором был утрачен/появился интересующий признак, и (2) экспрессия этих генов должна существенно изменяться во время развития интересующего признака у выбранного модельного вида, имеющего данный признак.
Чтобы идентифицировать гены, удовлетворяющие первому требованию, авторы разработали оригинальный алгоритм и компьютерную программу, позволяющие находить ортологи (прямые гомологи) генов выбранного модельного вида, имеющего интересующий признак, у других видов, не опираясь на уже существующие базы данных ортологов (такие базы далеко не полны, а так же дают существенное число ложных предсказаний), а путем широкоформатного скриннинга геномов с использованием критериев наилучшей гомологиии и локальной геномной синтении. В результате искомые гены возникшие/исчезнувшие на эволюционном этапе, совпадающим с этапом появления/исчезновения интересующего признака, определяются как те, у которых ортологи присутствуют/отсутствуют во всех видах, имеющих интересующий признак, но отсутствуют/присутствуют во всех видах, не имеющих его.
Гены, удовлетворяющие второму требованию, находятся путем глубокого секвенирования РНК, выделенной на разных стадиях развития данного признака.
Финальный список интересующих генов находится как пересечение списков генов, удовлетворяющих указанным двум требованиям.
удовлетворяющих указанным двум требованиям. В качестве доказательства принципа авторы использовали разработанный метод для поиска генов, отсутствующих у современных амниот (рептилий, птиц, млекопитающих) не способных к регенерации конечностей, но присутствующих у хорошо восстанавливающих ампутированные конечности анамниот (рыб и земноводных), у которых эти гены участвуют в регенерации конечностей. В результате было идентифицировано 57 таких генов, 30 из которых начинают активно экспрессироваться после ампутации хвоста головастика выбранного модельного вида, лягушки Xenopus tropicalis, а 27, наоборот, резко снижают свою экспрессию. Для трех из этих генов, c-c motif chemokine 4, eotaxin-like и ранее неизвестный ген, названный авторами sod4, были продемонстрированы существенные роли в регенерации хвоста. Интересно, что последний ген оказался членом нового четвертого семейства Cu/Zn- супероксиддисмутаз, утраченных амниотами, SOD4, которое впервые было описано в настоящей работе. Как известно, супероксиддисмутазы производят из супероксидрадикала перекись водорода , которая служит сигнальным агентом во множестве внутри- и внеклеточных процессов, в том числе, играет важную роль в регуляции регенерации.
Работа была поддержана грантом РНФ 23-74-30005 и опубликована в журнале Biology Direct.
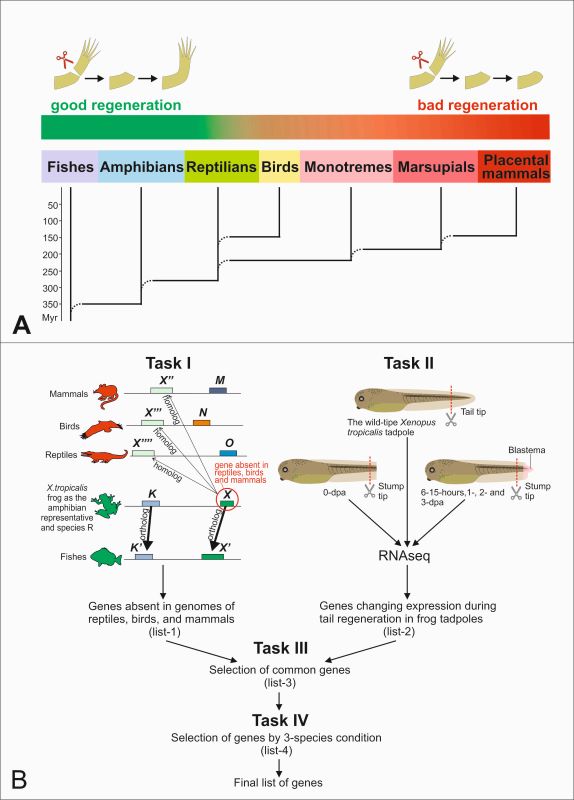
Рис. 1. Использование метода WINEGRET для поиска генов, регулирующих регенерацию придатков тела у рыб и земноводных, но отсутствующих у неспособных к регенерации рептилий, птиц и млекопитающих.
А. Снижение способности к регенерации крупных придатков тела, то есть конечностей и хвоста, у линии позвоночных от рыб и земноводных до плацентарных млекопитающих. Показаны большие группы позвоночных, возникшие последовательно в ходе эволюции.
Б. Схема метода WINEGRET на примере поиска генов, отсутствующих у млекопитающих, птиц и рептилий, но присутствующих у рыб и амфибий и участвующих в регенерации хвоста головастиков лягушки Xenopus tropicalis.
17 августа 2023 года


