Press-room / news / Science news /
Computer simulates body reaction to heat
For the first time, scientists at the M.M. Shemyakin and Yu. A. Ovchinnikov Institute of Bioorganic Chemistry of the Russian Academy of Sciences have successfully used computer generated simulations to study the TRPV1-Receptor temperature activation phenomenon for higher organisms. Up to this point, it was impossible to achieve such results in silico experiments due to imperfections in the computer simulation models. The results of the experiment were published in the Scientific Reports journal.
“We began this research study when our colleagues from the Laboratory of Neuroreceptors and Neuroregulators forwarded a request to study the dynamics and properties of the TRPV1-Receptor for the purposes of creating new anti-inflammatory and analgesic drugs,” explains Anton Chugunov, Candidate of Physical and Mathematical Sciences, collaborator at the Laboratory of Biomolecular Systems Simulations of the Institute of Bioorganic Chemistry of the Russian Academy of Sciences and one of the authors of the article, “Its former head and world-renown scientist, Yevgeny Grishin, was the initiator of this research. We have dedicated our work in his memory. We provided our colleagues with the necessary information relatively quickly, but during the course of the computations, discovered that we could observe something more – the effect of the receptor activation temperature. This is a very delicate process, and molecular dynamics simulation, which allows us to study the "behavior" of proteins, is still such an inaccurate ‘molecular microscope’ that we were amazed by the possibilities that opened before us.”
Belonging to the TRP-Receptor protein group, TRPV1 participates in the body’s temperature regulation system. Towards the end of the last century, an American scientist from the University of California in San Francisco, David Julius, for the first time unveiled the thermoregulatory function of this ion channel, which, when temperature rises, starts to pass sodium, magnesium and calcium ions through the cell membrane, while at the same time sends a signal to the cell to change the surrounding conditions. Later, Professor Julius decoded the TRPV1-Receptor spatial structure using the cryo-electron microscopy method, which allows detecting a small-sized object (10-15 nm) that is frozen under high-pressure. The resulting structure was defective in that the statically frozen receptor partially made it impossible to decode any dynamic changes, in which the scientists are very interested. Therefore, to date, the receptor activation process remains somewhat unclear. Anton, together with colleagues from the Institute of Bioorganic Chemistry, managed to fill this gap.
“We took two states of the receptor’s computer model: open and closed,” says Anton Chugunov. “To see how it worked, we carried out molecular dynamics calculations (which were quite impressive, lasting a year!), and also used the pore ‘computer-mapping’ method. We "unwrapped" the pore, that is, projected the properties of its internal channel-forming surface onto a plane, and clearly visualized them at different temperatures. Using a specially developed program, we monitored the changes in pore radius, which took place gradually over time, step by step, as happens in a living cell. The strong discontinuities that we registered on the graph indicated that the pore had changed its state from closed to open, but not vice versa. We knew that the receptor is activated at temperatures above 43 ° C, and anyone can feel this by placing his/her hand in hot water. We were therefore able to figure out the molecular dynamics for each state at four temperatures: two higher, and two lower than 43 ° C. The results were surprising.”
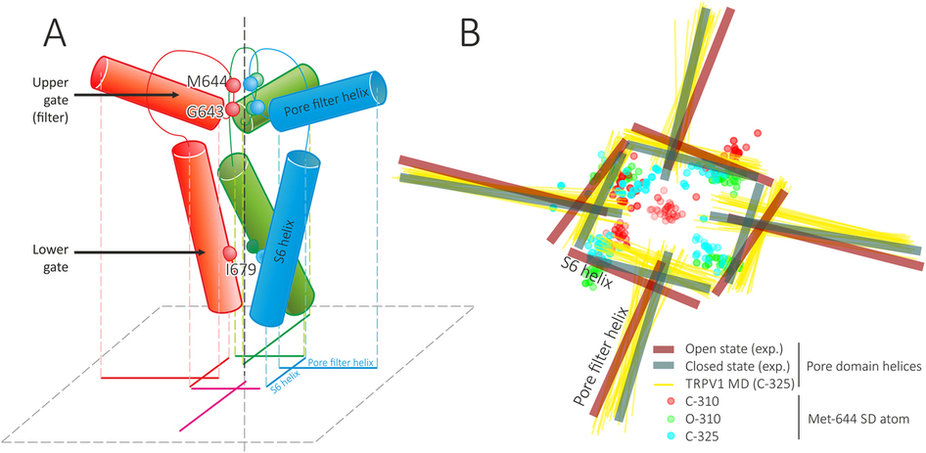
(А) Schematic representation of pore domain architecture and a projection that is used in panel (B) For each TRPV1 subunit (individually colored; the nearby one is omitted for clarity) two helices (pore filter (PF) and S6) and pore loop are shown.Spheres depict positions of several important residues: Gly 643 and Met 644 (upper gate) and Ile 679 (lower gate). A gunsight-like symbol is a projection of these eight helices (two from each subunit) on a membrane-parallel plane (dashed gray); two from one monomer are subscribed. Pore axis is shown with vertical dashed line. (B) Dynamic features of pore helices and Met 644 side chain in MD simulations. Thick lines show positions of TRPV1 pore helices (PF and S6) in closed (teal) and open (maroon) experimental structures. Thin yellow lines show positions of pore helices in C-325 MD trajectory. Circles depict projections of Met 644 SD atom in C-310 (red), O-310 (green) and C-325 (cyan) trajectories. Note that Met 644 SD atom from one of subunits blocks the pore in C-310 trajectory.
We logically assumed that the receptor for which the molecular dynamics calculation had been launched under the closed channel pore must remain closed at the two lower temperatures and open at the two higher ones, and vice versa. However, the computational experiment showed that in the closed position, the channel pore opened at high temperatures, but during the calculation of the pore’s open state, lower temperatures failed to close the pore for the duration of the computational time (up to 1 microsecond).
“The result that it is much easier to open the ion channel than close it was a kind of unbalance,” concludes Anton.
The data published show that modern computing capabilities help in investigating the ion channels and their operating characteristics in time intervals that are still impossible to detect experimentally. We are also able to observe the transitions within the receptor molecule as if through a ‘molecular microscope’.
This work is ultimately designed to soon establish a virtual TRPV1-Receptor that is suitable for predicting the structure of selective ligands. We have reason to expect that such ligands could find application in medicine as an analgesic medication. In this case, they would eliminate most of the side effects that are peculiar to modern medicines of the nonsteroidal anti-inflammatory drug groups.
september 13, 2016

