Пресс-центр / новости / Наука /
«Транскриптомика микобактерий как способ изучения механизмов контроля латентного туберкулеза»
До сих пор у ученых нет полного понимания того, что именно происходит в клетках M. tuberculosis в покоящейся стадии. Тем не менее, ответ на этот вопрос является насущной необходимостью, поскольку без него невозможно найти способы победить туберкулез. Сотрудники лаборатории Структуры и функции генов человека Института биоорганической химии имени академиков М. М. Шемякина и Ю. А. Овчинникова Т. Л. Ажикина и Д. В. Игнатов исследовали транскриптом представителей рода Mycobacterium и выявили возможные молекулярные механизмы, участвующие в установлении и поддержании состояния покоя бактериальных клеток. Результаты этой работы были представлены Д. В. Игнатовым на X Молодежной школе-конференции с международным участием «Актуальные аспекты современной микробиологии».
С бактерией Mycobacterium tuberculosis человечество знакомо уже около 5000 лет, однако до сих пор медицина не может полностью победить туберкулез – заболевание, которое вызывает этот микроорганизм. Дело в том, туберкулезная инфекция начинается с попадания M. tuberculosis в легкие человека. При этом быстро формируется иммунный ответ, и значительная часть бактерий поглощаются легочными макрофагами. Однако, поскольку микроорганизмы обладают способностью выживать внутри макрофага, часть бактерий остается целыми и невредимыми. Таким образом, дело не доходит до полного уничтожения возбудителей инфекции, и она переходит в скрытую форму.
Макрофаги, в свою очередь, могут подавлять рост бактерий, делая среду их обитания более кислой, а этого M. tuberculosis весьма не любит. После этого инфицированные макрофаги мигрируют во внутренние ткани легкого, где вокруг них возникает гранулёма – комплекс из разных иммунных клеток. Такая гранулёма может существовать на протяжении нескольких лет – это и соответствует стадии латентной туберкулезной инфекции. Сами же бактерии внутри гранулёмы переходят в дормантное состояние, во время которого они практически не делятся, а уровень их метаболизма достаточно низок. Однако, находясь в подобном состоянии, они становятся мало восприимчивы к воздействию многих внешних факторов – в том числе и лекарств.
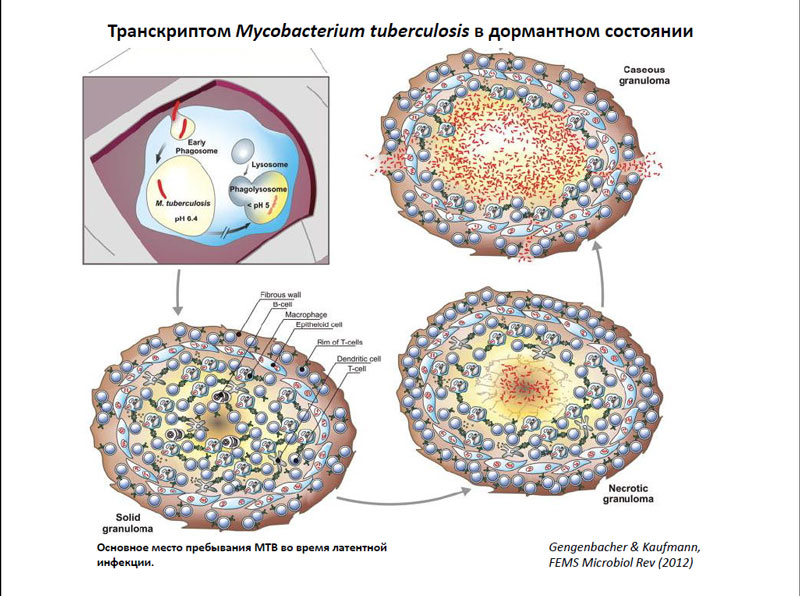
Транскриптом Mycobacterium tuberculosis в дормантном состоянии
Ответить на вопрос, что именно происходит в клетках M. tuberculosis внутри гранулёмы, достаточно сложно, поскольку нет модельных животных, на которых можно точно воспроизвести гранулёму, да и количество бактериальных клеток внутри нее весьма мало. Тем не менее, ответ на этот вопрос является насущной необходимостью, поскольку без него ученые так и не смогут найти способы победить туберкулез. Впрочем, сейчас разработаны различные in vitro модели, используя которые, можно перевести бактерий в дормантное состояние для того, чтобы изучить то, что происходит в это время внутри клеток M. tuberculosis.
Именно такое исследование и было проведено сотрудниками лаборатории Структуры и функции генов человека ИБХ РАН Т. Л. Ажикиной и Д. В. Игнатовым, которые изучали транскриптомику представителей рода Mycobacterium совместно с научными коллективами из Федерального исследовательского центра «Фундаментальные основы биотехнологии» РАН и Центрального научно-исследовательского института туберкулёза РАМН (оба - Москва). В процессе работы были выяснены многие интересные моменты, связанные с транскриптомом микобактерий.
Что же такое транскриптом? Это набор всех РНК, присутствующих в бактериальной клетке. Интересно, что более 80 процентов транскриптома составляют рибосомальные РНК. Доля же мРНК, хотя она и может варьировать в зависимости от фазы роста, как правило, невелика – менее 5 процентов. Все остальное – это транспортные (10 процентов) и различные регуляторные РНК. Основными типами регуляторных РНК у бактерий являются рибопереключатели и малые РНК. Рибопереключатели располагаются в нетранслируемых областях мРНК генов. Они работают следующим образом – при связывании с ними каких-либо низкомолекулярных веществ начинается активация или ингибирование трансляции этих самых генов. В отличие от рибопереключателей, малые РНК транскрибируются независимо от регулируемых ими генов. Комплементарное взаимодействие малых РНК с мРНК приводят либо к активации, либо к ингибированию трансляции этих генов.
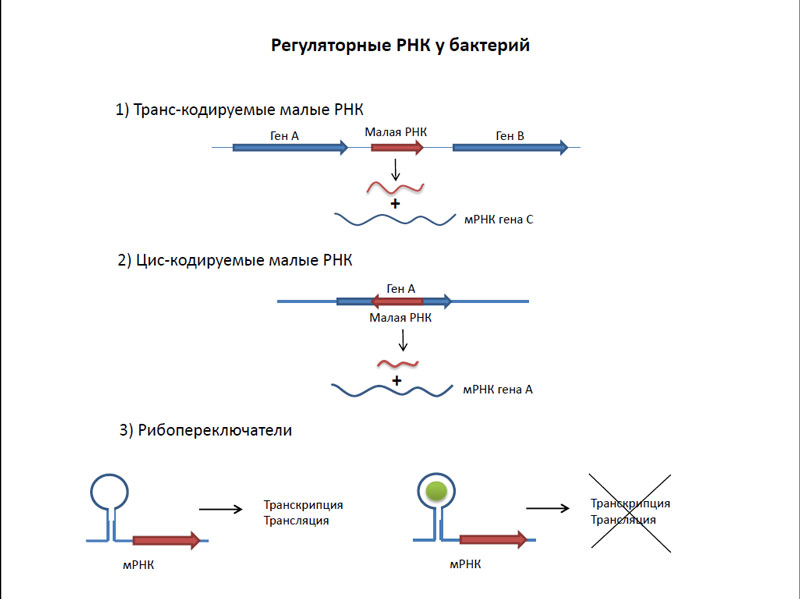
Регуляторные РНК у бактерий
Таким образом, изучая транскриптом, можно узнать, какие именно гены активны в клетке в данный момент, а какие, наоборот, не работают. Также подобные исследования дают представление о том, как регулируются многие процессы, происходящие в клетках микроорганизмов. Это же, в свою очередь, позволит понять, как именно клетка микобактерии переходит в дормантное состояние и что происходит с ней в этой стадии жизненного цикла.
При изучении транскриптома микобактерий исследователи использовали RNA-seq, или метод секвенирования РНК. В его основе лежит секвенирование фрагментов кДНК с помощью секвенаторов нового поколения. С помощью метода RNA-seq можно производить поиск новых генов у бактерий, включая гены регуляторных и некодирующих РНК, с предельной точностью (до одного нуклеотида) картировать границу транскриптов, измерять уровни транскрипции генов в широком диапазоне и определять структуры оперонов.
Суть этой методики заключается в следующем - сначала из бактериальной клетки выделялась вся РНК. Потом нужно было избавиться от самой многочисленной, но совершенно не интересной для нашего исследования рибосомальной РНК. Затем остаток фрагментировался и к каждому фрагменту прикреплялись 5’ и 3’ концевые адаптеры, которые нужны для затравки синтеза кДНК. После этого происходил синтез и амплификация кДНК, а потом – секвенирование этих ДНК с помощью секвенаторов нового поколения. На основе полученных в результате секвенирования данных строились карты генов бактерий, которые анализировались при помощи методов биоинформатики.
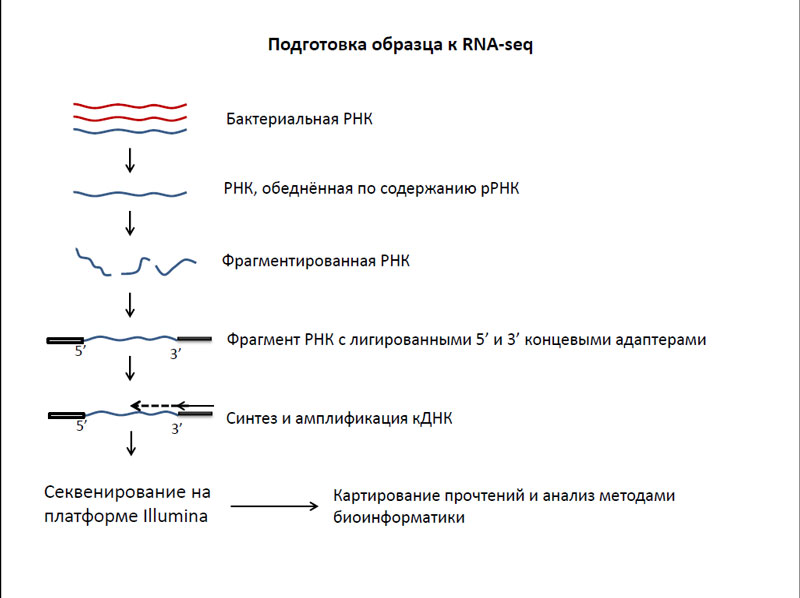
Подготовка образца к RNA-seq
Объектами исследования были клетки на Mycobacterium avium и Mycobacterium tuberculosis – последние при этом находились в дормантном состоянии. По словам Д. В. Игнатова, «когда мы начинали нашу работу, то существовало только одно подобное исследование, проводившееся на клетках M. tuberculosis в культуре. Сейчас же имеется около десятка работ, посвященных изучению транскриптомов этих бактерий». Это говорит о том, что до сих пор процессы, происходящие в клетках микобактерий в покоящейся стадии, изучены достаточно плохо.
Заражение патогенным микроорганизмом Mycobacterium avium приводит к возникновению целого ряда болезней у животных и человека. Так, подвиды M. avium avium и M. avium silvaticum вызывают заболевания птиц, а подвид M. avium paratuberculosis – болезнь Джонса жвачных животных. Для человека наибольшую опасность представляет подвид M. avium hominissuis, который широко распространен в окружающей среде, который при попадании в человеческий организм способен вызвать диссеминированные заболевания легких у людей с иммунодефицитом. Следует заметить, что патогенез Mycobacterium avium схож с таковым M. tuberculosis – этот микроорганизм также способен концентрироваться внутри макрофагов. Поэтому изучение транскриптомов данного микроорганизма может помочь понять, какие процессы происходят в дормантных клетках M. tuberculosis, которые сложно изучать in vivo.
После секвенирования транскриптома Mycobacterium avium было получено несколько десятков миллионов прочтений, причем часть из них оказалась локусами рибосомальных РНК – и это несмотря на то, что ее старались удалить. Часть локусов картировалась в области мРНК, часть – в межгенных областях (причем таковых было даже больше, чем транслируемых локусах), и часть из них оказалась антисмысловыми последовательностями частей мРНК. Очевидно, что два последних типа прочтения имеют отношения к регуляторным малым РНК.
В итоге исследователям удалось идентифицировать несколько малых РНК из транскриптома M. avium и сравнить их с таковыми M. tuberculosis. Выяснилась, что большая часть этих малых РНК имеет гомологи у M. tuberculosis за исключением нескольких из них. Так, в геноме M. avium отсутствуют гомологи двух малых РНК M. tuberculosis, такие как MTS0479 (предположительно работает тогда, когда клетка испытывает кислотный стресс) и MTS1338 (синтезируется во время стационарной фазы и при гипоксии). В свою очередь у M. tuberculosis отсутствуют гомологи четырех малых РНК M. avium, функции которых не известны.
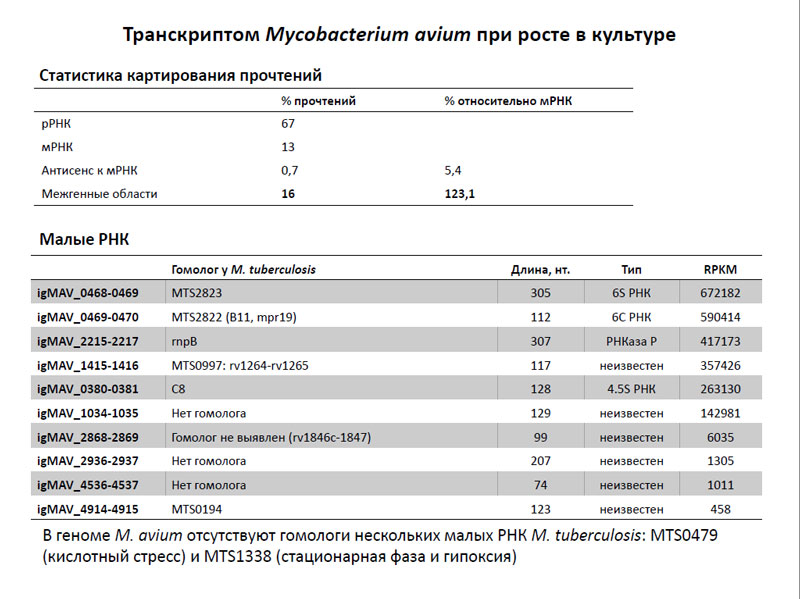
Транскриптом Mycobacterium avium при росте в культуре
Вторым источником прочтений были локусы из 5’-нетранслируемых областей. Исследователи разработали алгоритм картирования транскрипционных старт-сайтов (ТСС) на транскрипционном профиле. И тут выяснилась интересная вещь - 33% ТСС совпадали со старт-кодонами, то есть они соответствовали безлидерным мРНК. Это весьма необычно для бактерий, у которых такие мРНК встречаются крайне редко – они известны всего лишь для нескольких десятков генов. Совершенно непонятно, почему их было так много у микобактерий – по мнению авторов работы, этот вопрос требует дальнейшего детального изучения. Несколько же достаточно длинных 5’-нетранслируемых областей соответствовали рибопереключателям различных генов.
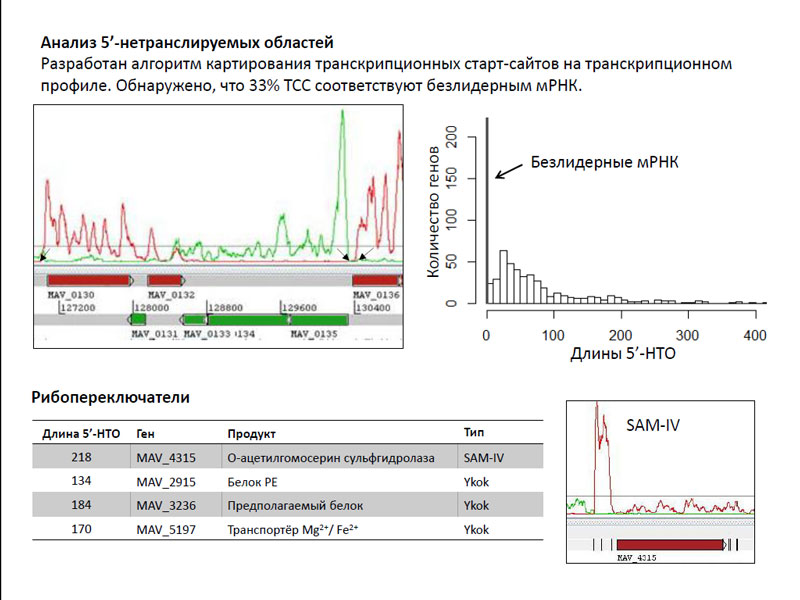
Анализ 5’-нетранслируемых областей
Исследования транскриптомов Mycobacterium tuberculosis в дормантном состоянии проводились на модели, которая была разработана в лаборатории проф. А.С. Капрельянца из Института биохимии им. А.Н. Баха РАН. Клетки M. tuberculosis культивировались в жидкой среде, где отсутствовали ионы калия. Выращенные таким образом бактерии приобретали весьма интересные свойства: они утрачивали способность расти на твердой среде, и были нечувствительны к воздействию нескольких разновидностей антибиотиков. Кроме того, эти микроорганизмы имели необычную морфологию клеток, у них был понижен синтез АТФ и дыхание, а также было понижено содержание липидов. Что касается уровня транскрипции, то он тоже был достаточно низким. Через два дня такой культивации в колонию M. tuberculosis добавляли антибиотик рифампицин в ингибирующей концентрации (5 μg/ml), который убивал все активно делящиеся клетки. Таким образом, в культуре оставались только дормантные микроорганизмы.
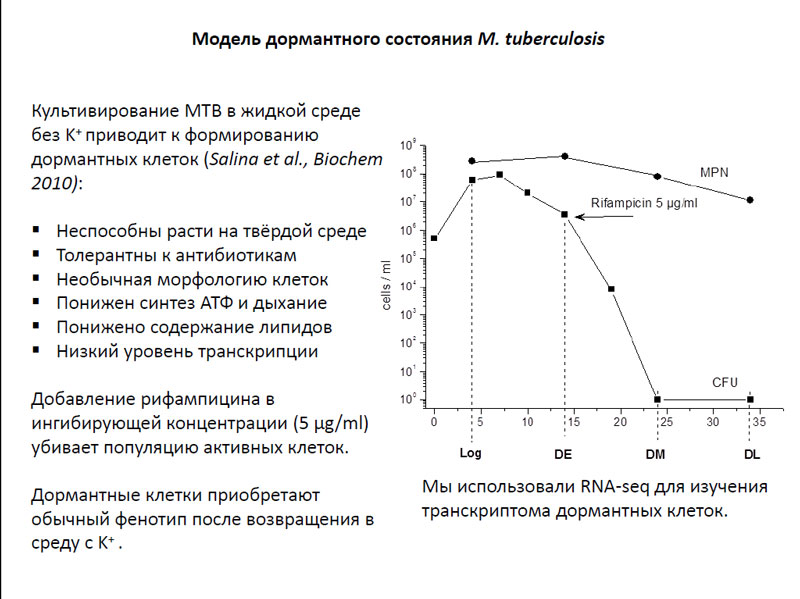
Модель дормантного состояния M. tuberculosis
Т. Л. Ажикина и Д.В. Игнатов изучили транскриптомику клеток M. tuberculosis на ранней дормантной стадии, то есть перед добавлением рифампицина, а также тех, что находились на стадии позднего дормантного состояния, которое наступило через несколько дней после инъекции антибиотка. Интересным свойством транскриптома дормантных клеток было разрезание 23S рибосомальной РНК. Исследователям удалось картировать точку разрезания 23S рРНК – выяснилось, что она располагается достаточно далеко от каталитического центра рибосомы, и, по-видимому, не влияет на синтез белка. Однако о том, какой агент вызывает это разрезание и оказывает ли разрезание на физиологию бактериальной клетки, ничего не известно. Тем не менее, не исключено, что этот механизм активно способствует переходу бактериальных клеток в дормантное состояние, а также поддерживает его.
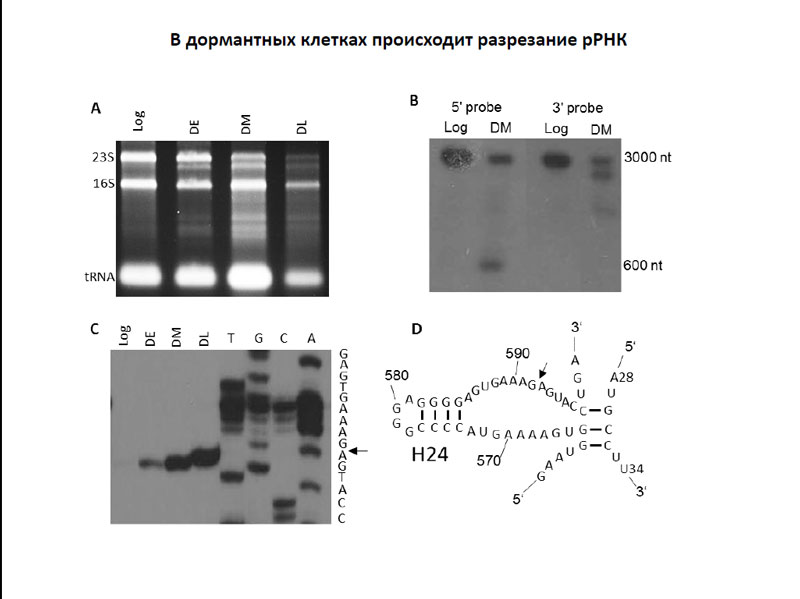
Разрезание рРНК в дормантных клетках
Кроме того, было установлено, что в дормантных клетках значительно понижается уровень транскрипции (особенно это было заметно в клетках на поздней дормантной стадии), а суммарное количество мРНК понижается в 30-50 раз по сравнению с ранней дормантной стадией. Тем не менее, несмотря на столь малое количество мРНК, исследователям удалось измерить уровень экспрессии белок-кодирующих генов. Авторы работы сконцентрировали свое внимание на тех генах, которые изменяют свою экспрессию при культивировании в среде без ионов калия, а также на тех, чья экспрессия меняется при добавлении рифампицина.
В результате выяснилось, что при культивировании в среде без ионов калия значительно понижается экспрессия генов, чьи белки участвуют в дыхании, а также генов субъединиц АТФ-синтетазы. Среди генов, чья активность повышалась при отсутствии ионов калия, следует отметить гены PE-PGRS, которых достаточно много у микобактерий, хотя до сих пор их функция совершенно непонятна. Известно лишь, что эти гены распространились по геному микобактерий относительно недавно и в первую очередь они характерны именно для патогенных микобактерий.
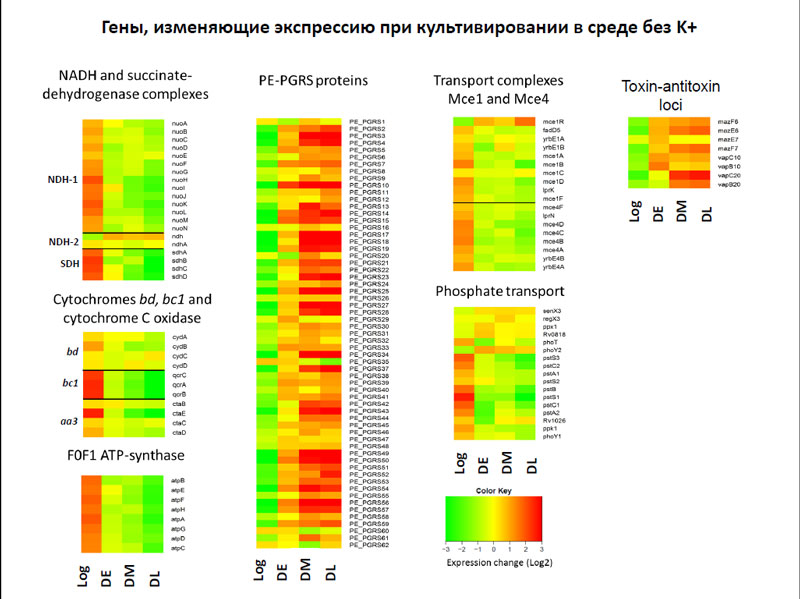
Гены, изменяющие экспрессию при культивировании в среде без K+
После же добавления рифампицина в клетках бактерий понижалась активность генов, кодирующих ферменты цикла трикарбоновых кислот и рибосомальные белки.
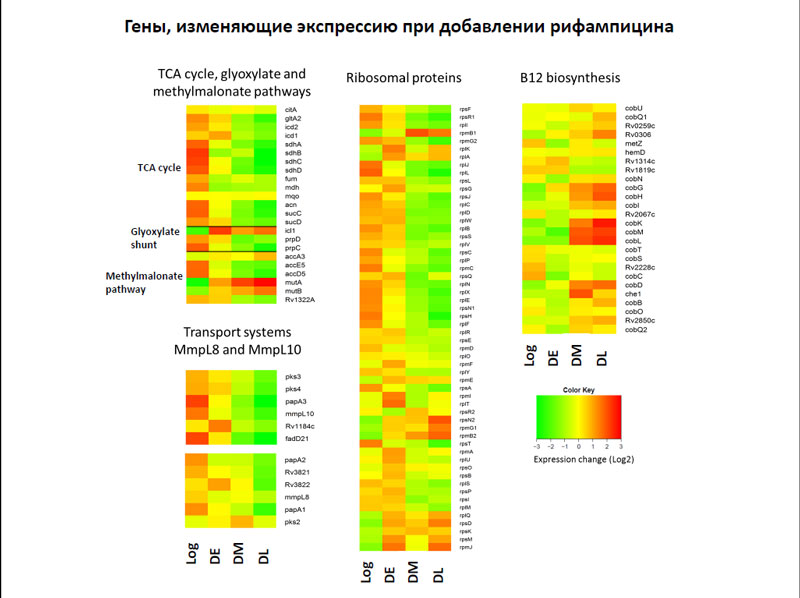
Гены, изменяющие экспрессию при добавлении рифампицина
Также была исследована экспрессия нескольких малых РНК в клетках, находящихся в раннем и позднем дормантных состояниях. Было отмечено, что уровень экспрессии таких малых РНК, MTS0997, MTS2823 и MTS1338 даже в покоящихся бактериях остается достаточно высоким. Интересно, что экспрессия MTS1338 контролируется транскрипционным фактором DosR, который регулирует гены, участвующие в переходе в дормантное состояние. Исходя из этих данных, исследователи предположили, что именно эта малая РНК может играть большую роль в установлении дормантного состояния.
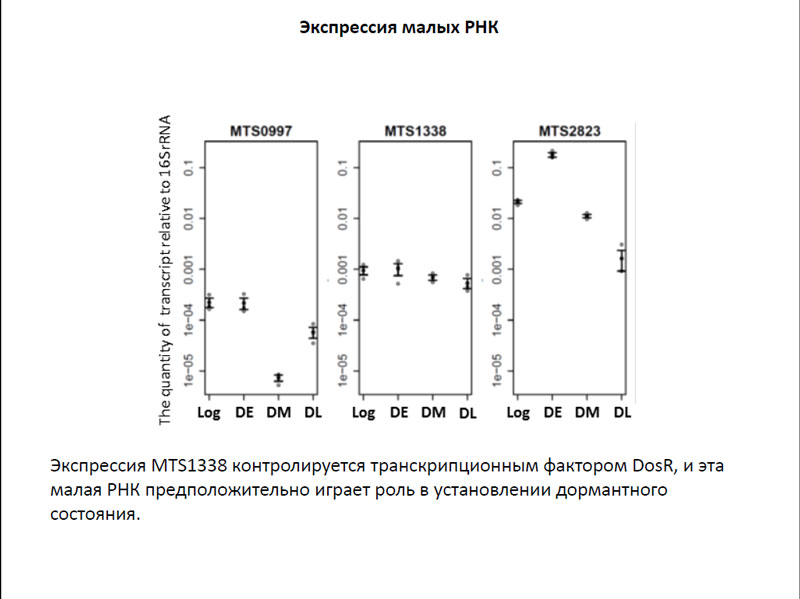
Экспрессия малых РНК
Таким образом, Т.Л. Ажикиной и Д.В. Игнатовым были впервые получены данные об изменениях, происходящих в транскриптоме Mycobacterium tuberculosis при переходя в дормантное состояние in vitro в условиях недостатка калия. Был также предложены возможные механизмы, участвующие в установлении и поддержании этого состояния покоя. Все это может помочь разработке способов борьбы с возбудителем туберкулеза, находящимся в покоящейся стадии.
Литература
1) Ignatov, D., Malakho, S., Majorov, K., Skvortsov, T., Apt, A. and Azhikina, T. (2013) RNA-Seq analysis of Mycobacterium avium non-coding transcriptome. PloS one, 8, e74209.
2) Ignatov D., Salina E., Fursov M., Skvortsov T., Azhikina T., Kaprelyants A. (2015). “Dormant non-culturable Mycobacterium tuberculosis retains stable low-abundant mRNA”. BMC Genomics, in press.
6 ноября 2015 года

